mavoglurant – Physiologically-based PK
Wenping Wang
2022-03-26
Source:vignettes/mavoglurant.Rmd
mavoglurant.Rmd
nlmixr
Building on the first simple example, we can be more ambitious, and try a full PBPK model. This one was published for mavoglurant (Wendling et al. 2016).
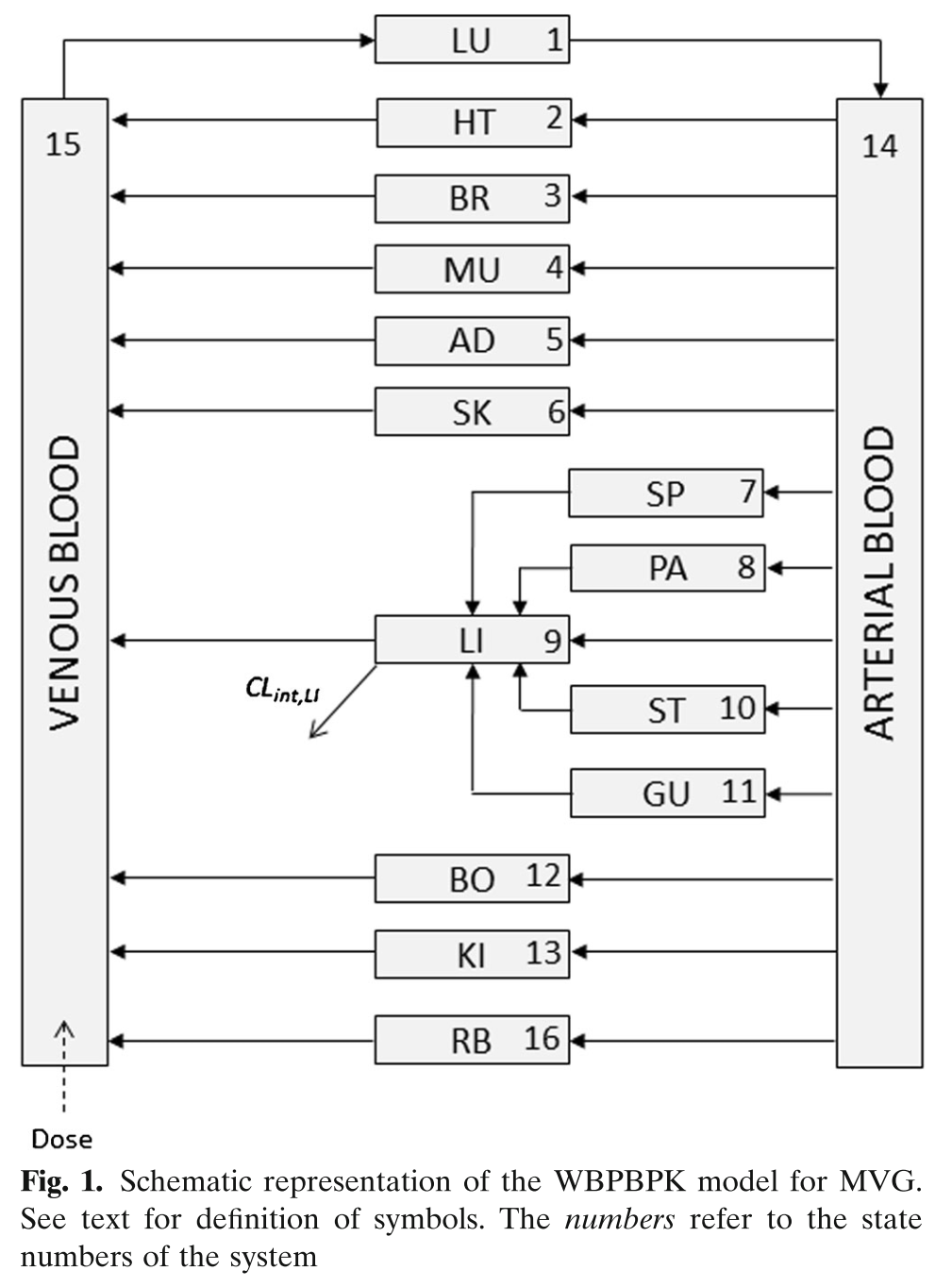
Model Schematic
nlmixr model
library(nlmixr)
library(xpose)
library(xpose.nlmixr)
library(ggplot2)
pbpk <- function(){
ini({
##theta=exp(c(1.1, .3, 2, 7.6, .003, .3))
lKbBR = 1.1
lKbMU = 0.3
lKbAD = 2
lCLint = 7.6
lKbBO = 0.03
lKbRB = 0.3
eta.LClint ~ 4
add.err <- 1
prop.err <- 10
})
model({
KbBR = exp(lKbBR)
KbMU = exp(lKbMU)
KbAD = exp(lKbAD)
CLint= exp(lCLint + eta.LClint)
KbBO = exp(lKbBO)
KbRB = exp(lKbRB)
## Regional blood flows
CO = (187.00*WT^0.81)*60/1000; # Cardiac output (L/h) from White et al (1968)
QHT = 4.0 *CO/100;
QBR = 12.0*CO/100;
QMU = 17.0*CO/100;
QAD = 5.0 *CO/100;
QSK = 5.0 *CO/100;
QSP = 3.0 *CO/100;
QPA = 1.0 *CO/100;
QLI = 25.5*CO/100;
QST = 1.0 *CO/100;
QGU = 14.0*CO/100;
QHA = QLI - (QSP + QPA + QST + QGU); # Hepatic artery blood flow
QBO = 5.0 *CO/100;
QKI = 19.0*CO/100;
QRB = CO - (QHT + QBR + QMU + QAD + QSK + QLI + QBO + QKI);
QLU = QHT + QBR + QMU + QAD + QSK + QLI + QBO + QKI + QRB;
## Organs' volumes = organs' weights / organs' density
VLU = (0.76 *WT/100)/1.051;
VHT = (0.47 *WT/100)/1.030;
VBR = (2.00 *WT/100)/1.036;
VMU = (40.00*WT/100)/1.041;
VAD = (21.42*WT/100)/0.916;
VSK = (3.71 *WT/100)/1.116;
VSP = (0.26 *WT/100)/1.054;
VPA = (0.14 *WT/100)/1.045;
VLI = (2.57 *WT/100)/1.040;
VST = (0.21 *WT/100)/1.050;
VGU = (1.44 *WT/100)/1.043;
VBO = (14.29*WT/100)/1.990;
VKI = (0.44 *WT/100)/1.050;
VAB = (2.81 *WT/100)/1.040;
VVB = (5.62 *WT/100)/1.040;
VRB = (3.86 *WT/100)/1.040;
## Fixed parameters
BP = 0.61; # Blood:plasma partition coefficient
fup = 0.028; # Fraction unbound in plasma
fub = fup/BP; # Fraction unbound in blood
KbLU = exp(0.8334);
KbHT = exp(1.1205);
KbSK = exp(-.5238);
KbSP = exp(0.3224);
KbPA = exp(0.3224);
KbLI = exp(1.7604);
KbST = exp(0.3224);
KbGU = exp(1.2026);
KbKI = exp(1.3171);
##-----------------------------------------
S15 = VVB*BP/1000;
C15 = Venous_Blood/S15
##-----------------------------------------
d/dt(Lungs) = QLU*(Venous_Blood/VVB - Lungs/KbLU/VLU);
d/dt(Heart) = QHT*(Arterial_Blood/VAB - Heart/KbHT/VHT);
d/dt(Brain) = QBR*(Arterial_Blood/VAB - Brain/KbBR/VBR);
d/dt(Muscles) = QMU*(Arterial_Blood/VAB - Muscles/KbMU/VMU);
d/dt(Adipose) = QAD*(Arterial_Blood/VAB - Adipose/KbAD/VAD);
d/dt(Skin) = QSK*(Arterial_Blood/VAB - Skin/KbSK/VSK);
d/dt(Spleen) = QSP*(Arterial_Blood/VAB - Spleen/KbSP/VSP);
d/dt(Pancreas) = QPA*(Arterial_Blood/VAB - Pancreas/KbPA/VPA);
d/dt(Liver) = QHA*Arterial_Blood/VAB + QSP*Spleen/KbSP/VSP + QPA*Pancreas/KbPA/VPA + QST*Stomach/KbST/VST + QGU*Gut/KbGU/VGU - CLint*fub*Liver/KbLI/VLI - QLI*Liver/KbLI/VLI;
d/dt(Stomach) = QST*(Arterial_Blood/VAB - Stomach/KbST/VST);
d/dt(Gut) = QGU*(Arterial_Blood/VAB - Gut/KbGU/VGU);
d/dt(Bones) = QBO*(Arterial_Blood/VAB - Bones/KbBO/VBO);
d/dt(Kidneys) = QKI*(Arterial_Blood/VAB - Kidneys/KbKI/VKI);
d/dt(Arterial_Blood) = QLU*(Lungs/KbLU/VLU - Arterial_Blood/VAB);
d/dt(Venous_Blood) = QHT*Heart/KbHT/VHT + QBR*Brain/KbBR/VBR + QMU*Muscles/KbMU/VMU + QAD*Adipose/KbAD/VAD + QSK*Skin/KbSK/VSK + QLI*Liver/KbLI/VLI + QBO*Bones/KbBO/VBO + QKI*Kidneys/KbKI/VKI + QRB*Rest_of_Body/KbRB/VRB - QLU*Venous_Blood/VVB;
d/dt(Rest_of_Body) = QRB*(Arterial_Blood/VAB - Rest_of_Body/KbRB/VRB);
C15 ~ add(add.err) + prop(prop.err)
})
}
dat = read.csv("Mavoglurant_A2121_nmpk.csv")
dat$occ = unlist(with(dat, tapply(EVID, ID, function(x) cumsum(x>0))))
dat = subset(dat, occ==1)
dat = subset(dat, ID<812) ## First 20
dat = subset(dat, EVID>0 | DV>0)
dat$CMT[dat$CMT == 0] <- 1;
dat$CMT[dat$EVID == 1] <- "Venous_Blood" ## Compartment dosed to is Venous Blood
dat$CMT[dat$EVID != 1] <- "C15" ## Observing C15
gofs <- function(fit){
################################################################################
## Standard plots
################################################################################
plot(fit);
xpdb <- xpose_data_nlmixr(fit) ## Convert to nlmixr object
print(dv_vs_pred(xpdb) +
ylab("Observed Mavoglurant Concentrations (ng/mL)") +
xlab("Population Predicted Mavoglurant Concentrations (ng/mL)"));
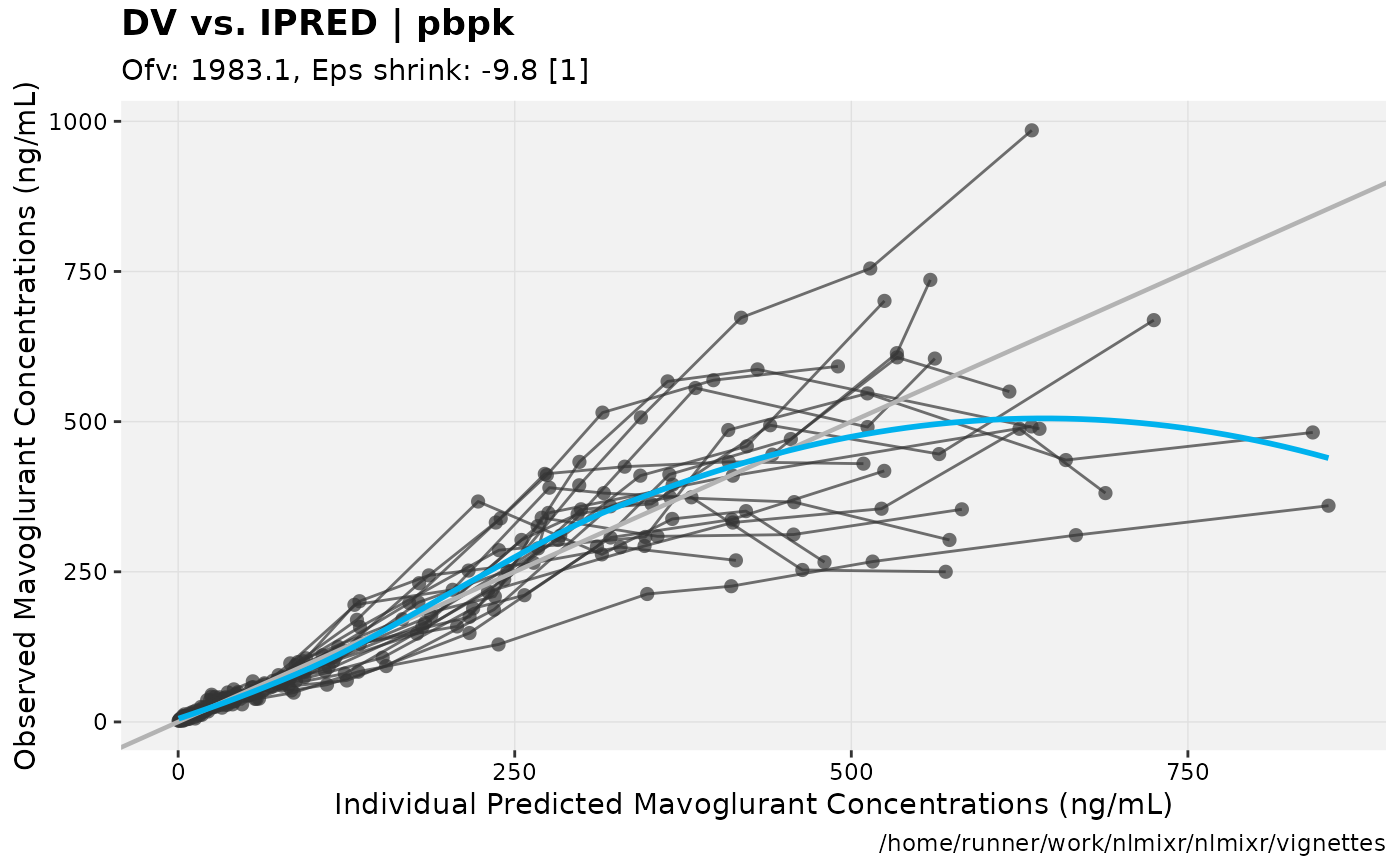
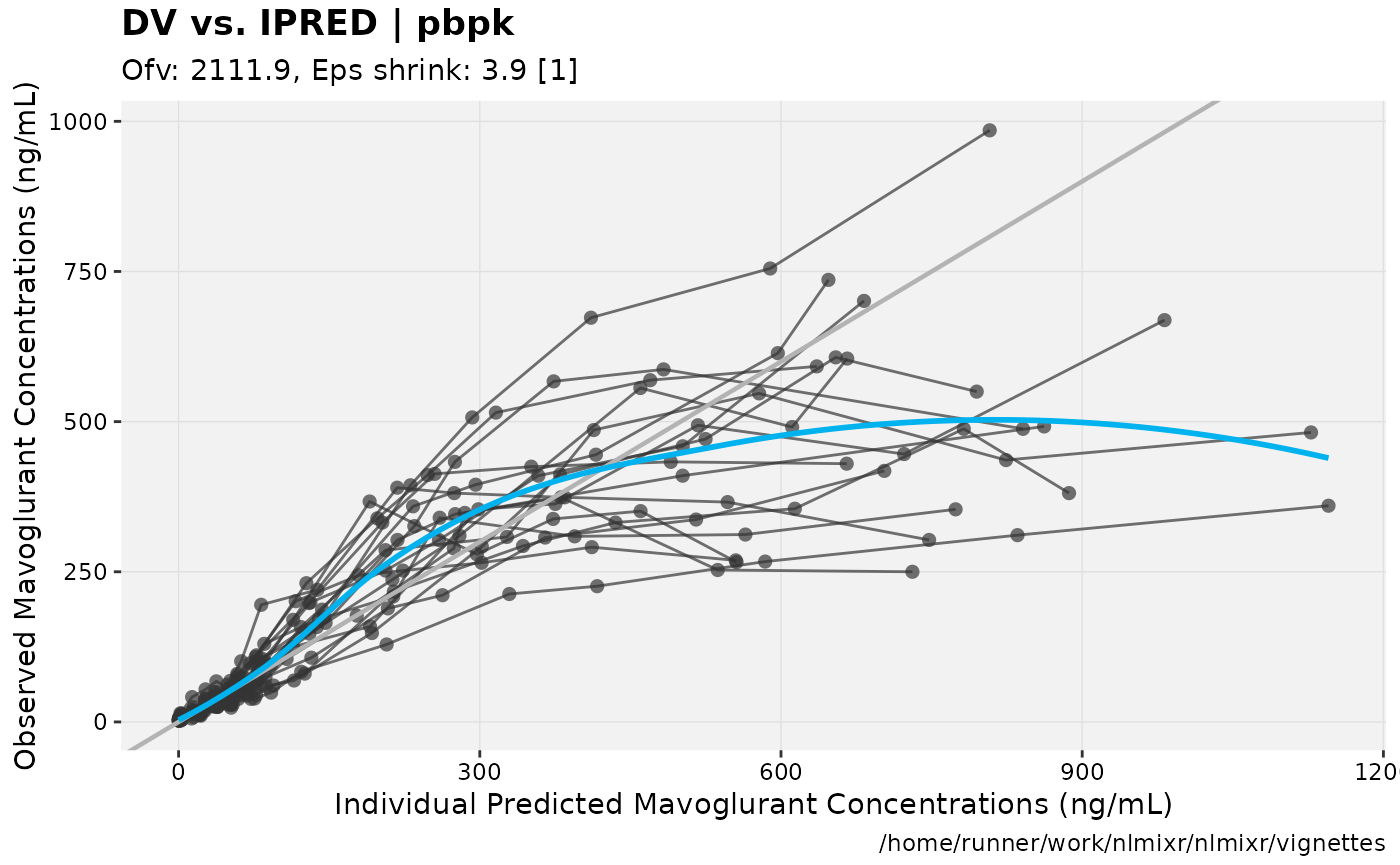
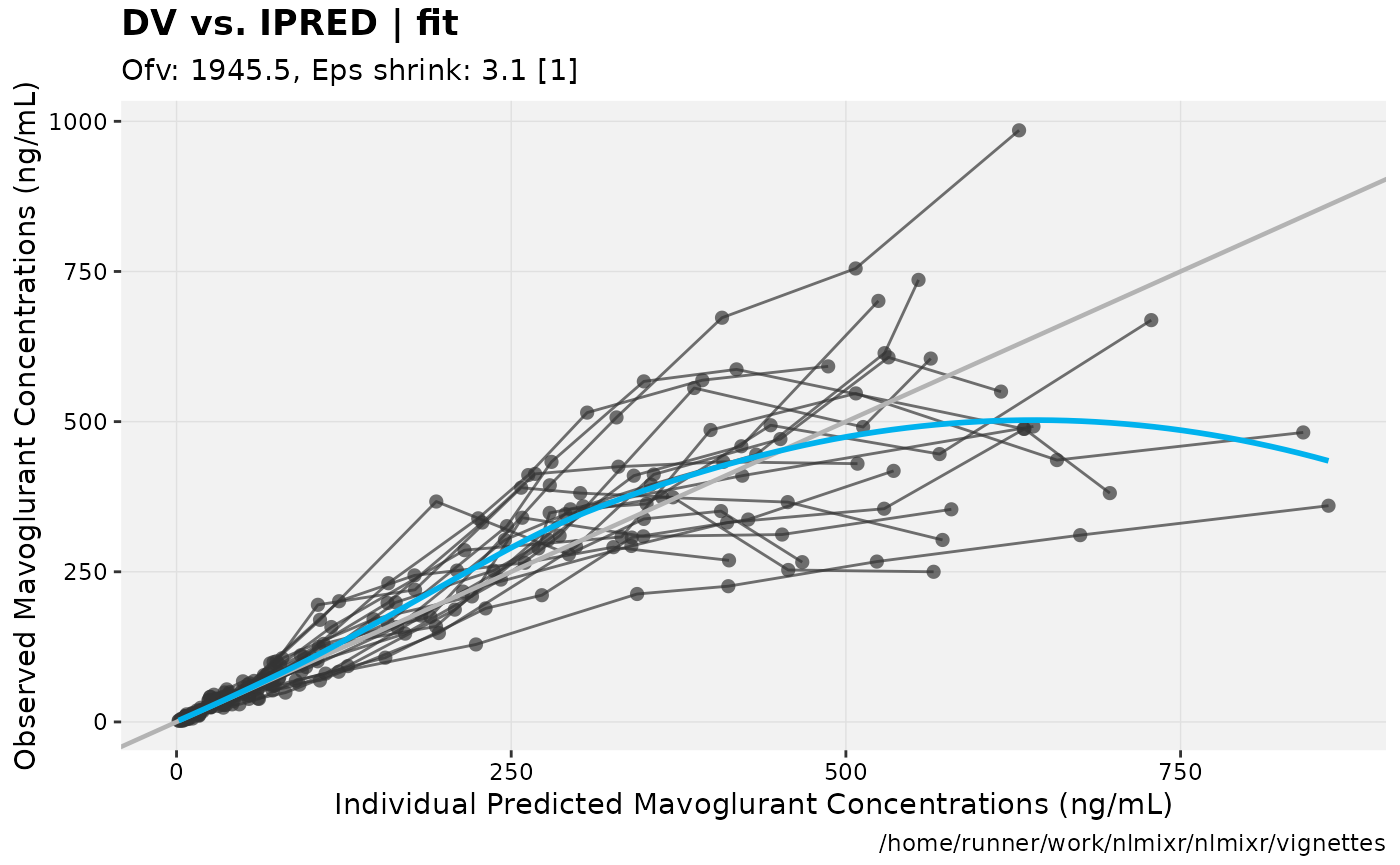
print(dv_vs_ipred(xpdb) +
ylab("Observed Mavoglurant Concentrations (ng/mL)") +
xlab("Individual Predicted Mavoglurant Concentrations (ng/mL)"));
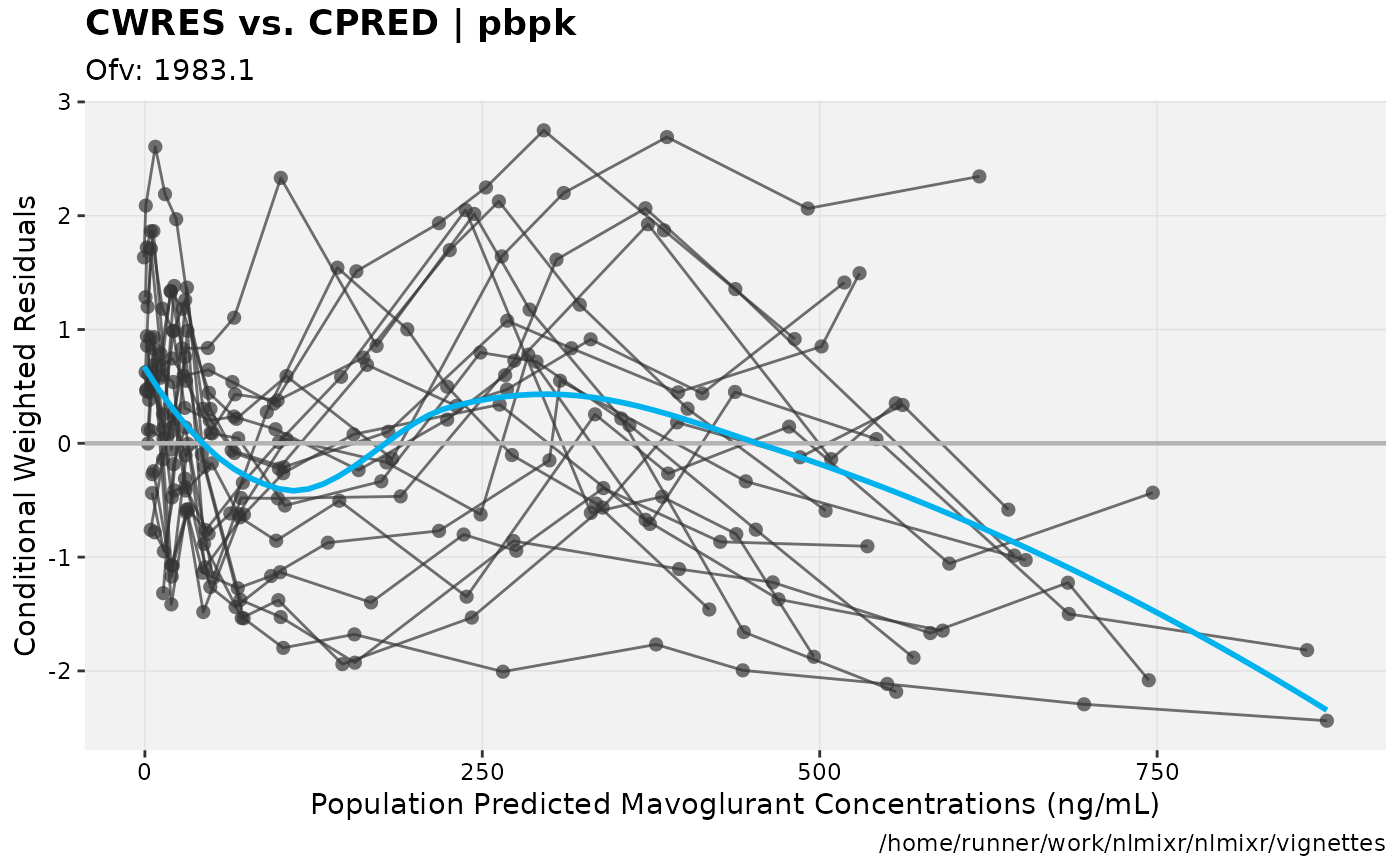
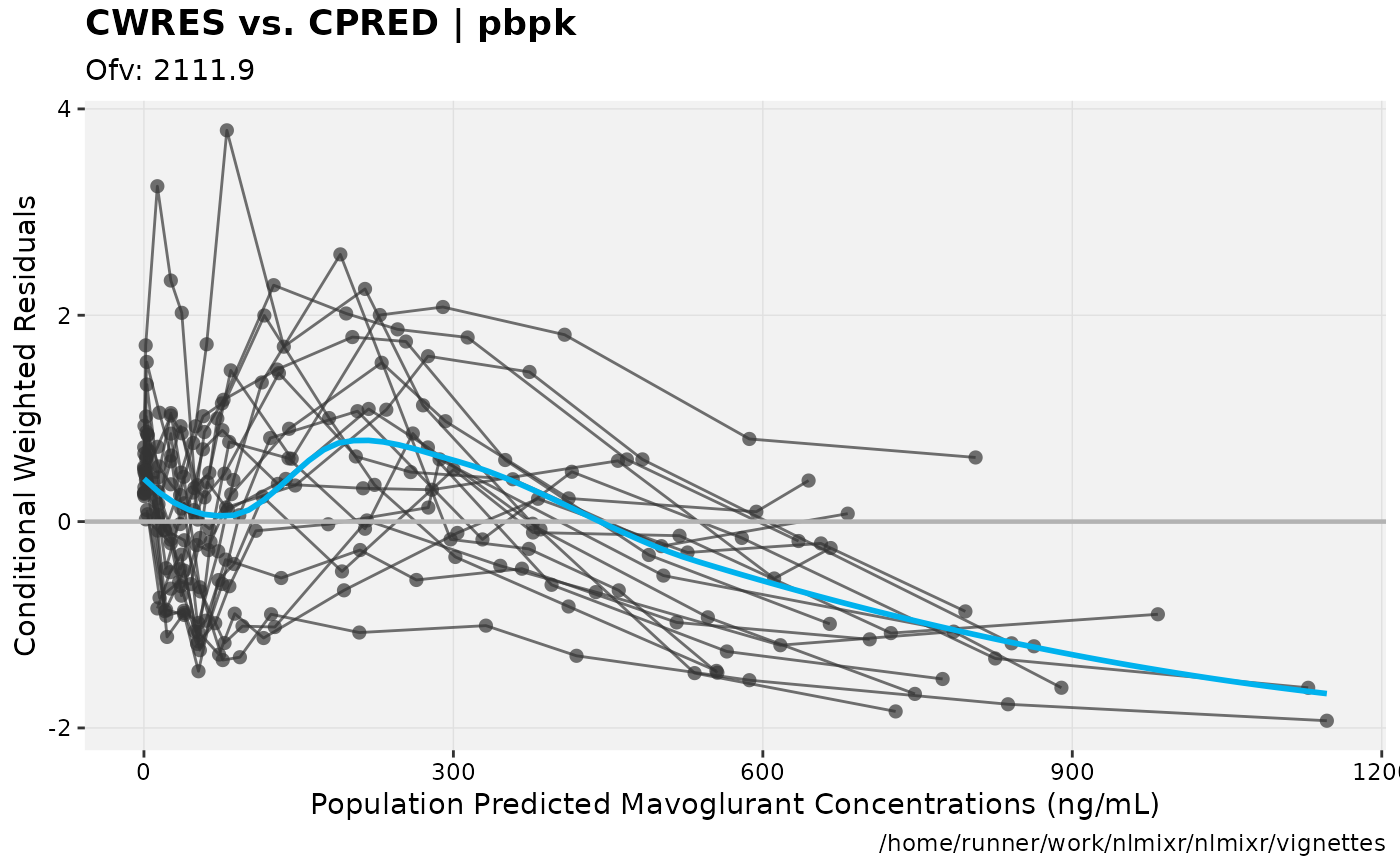
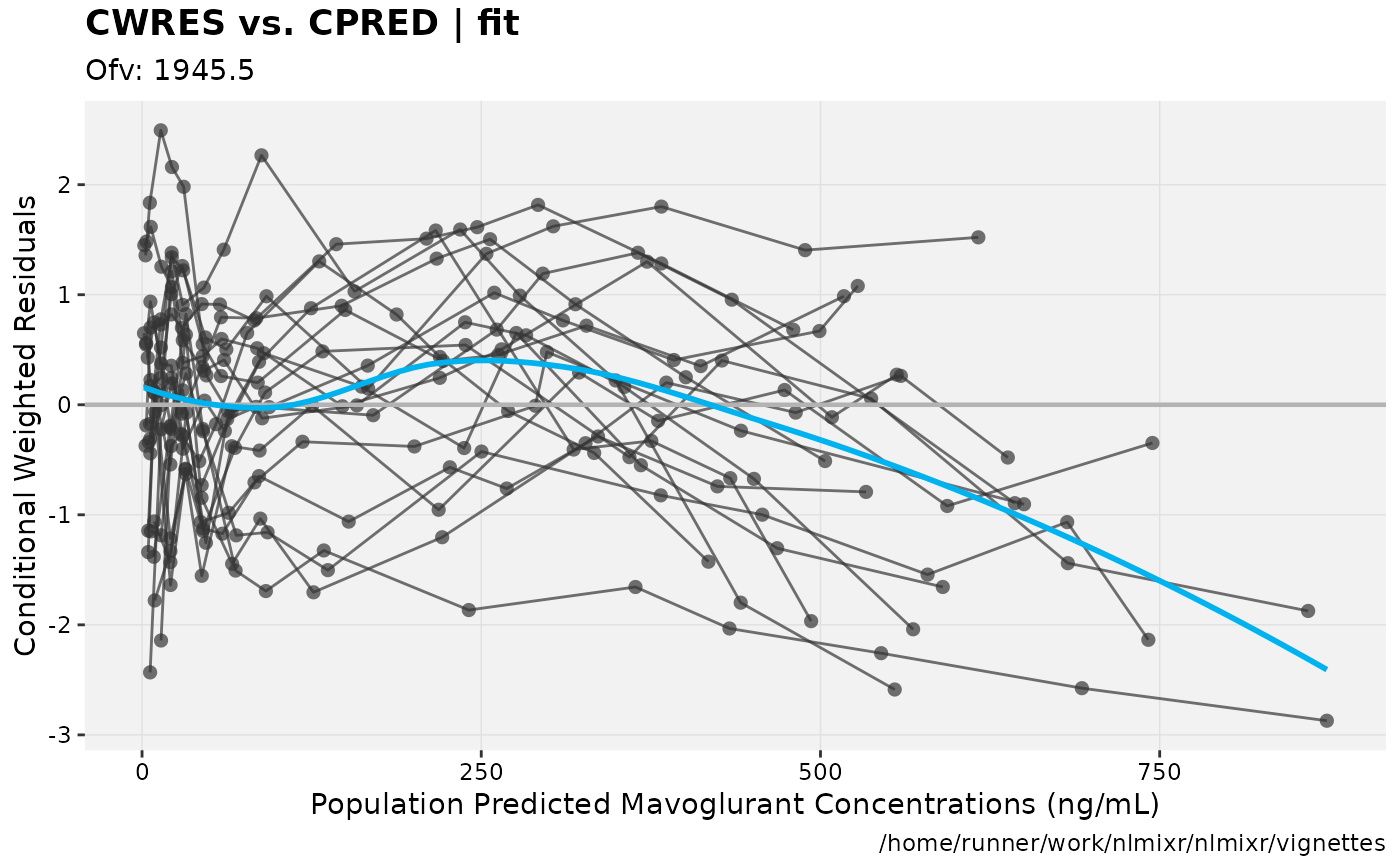
print(res_vs_pred(xpdb) +
ylab("Conditional Weighted Residuals") +
xlab("Population Predicted Mavoglurant Concentrations (ng/mL)"));
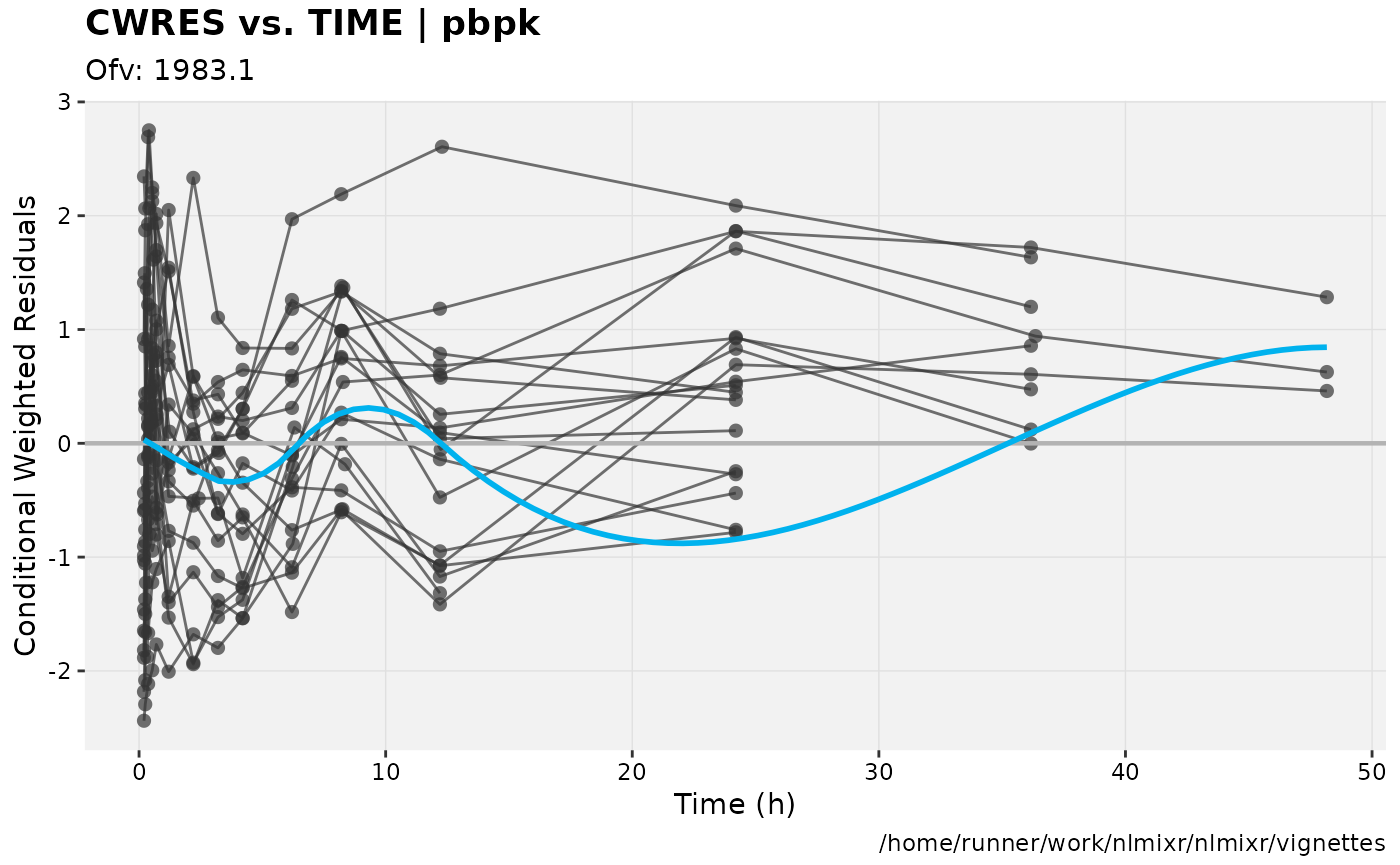
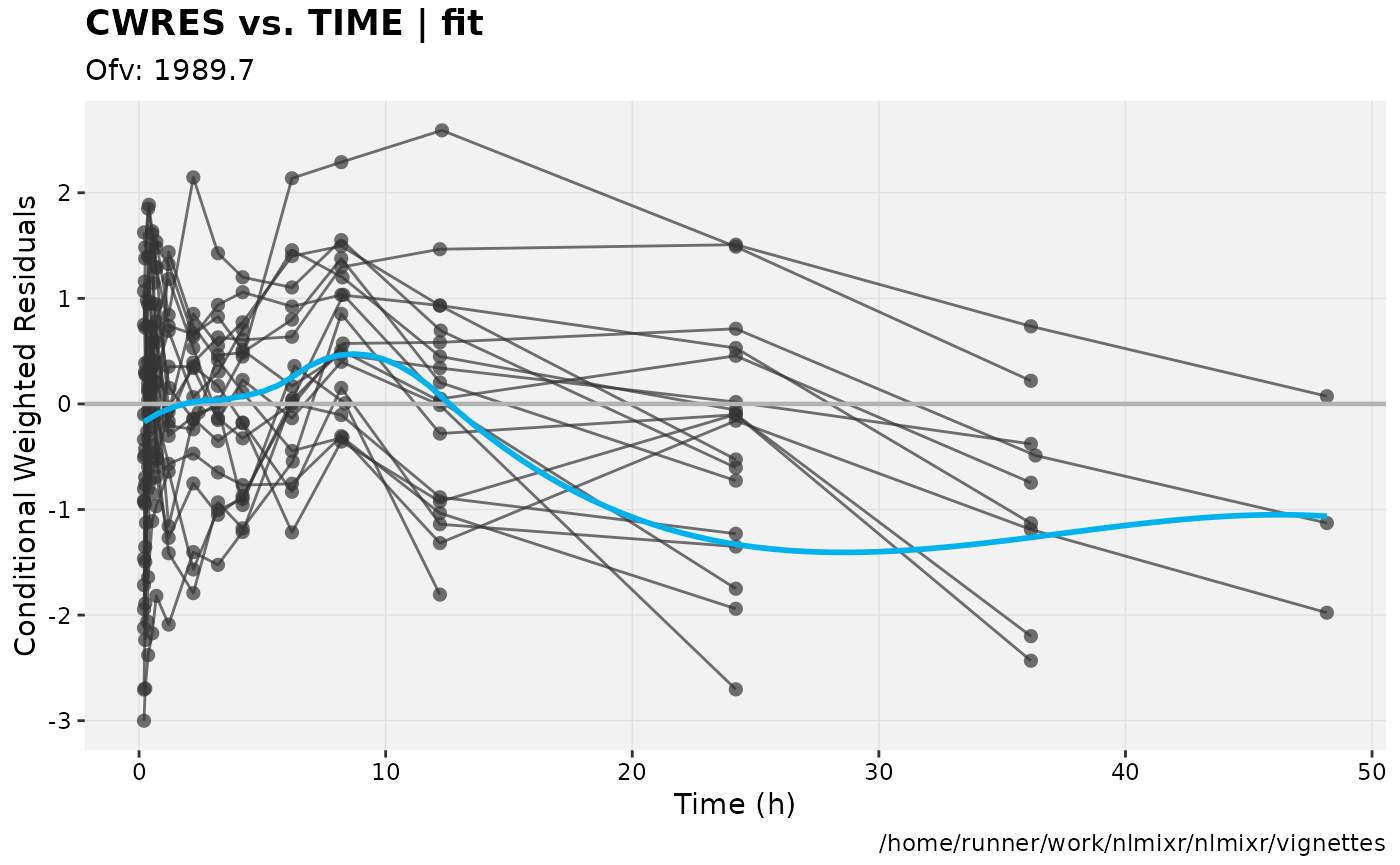
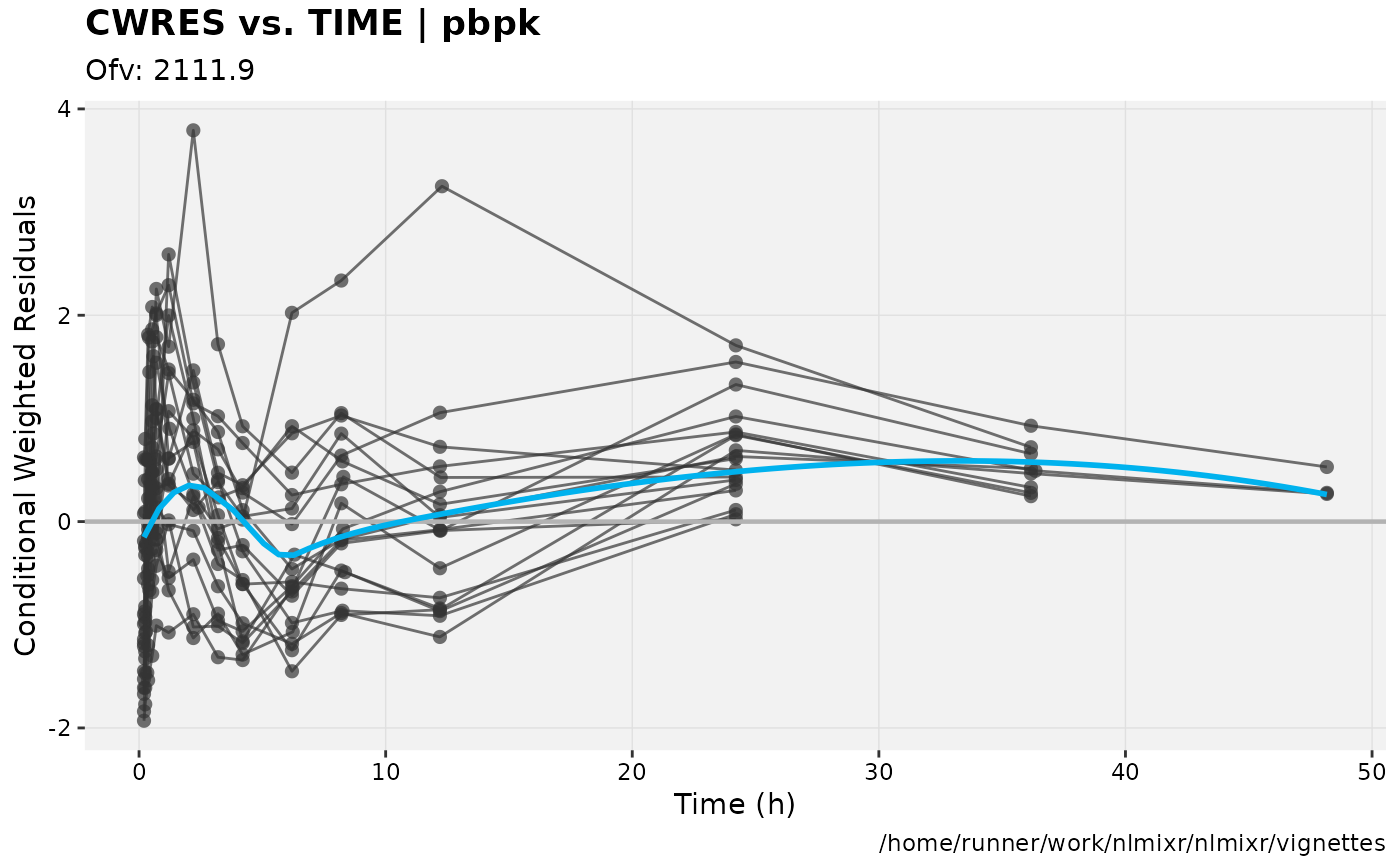
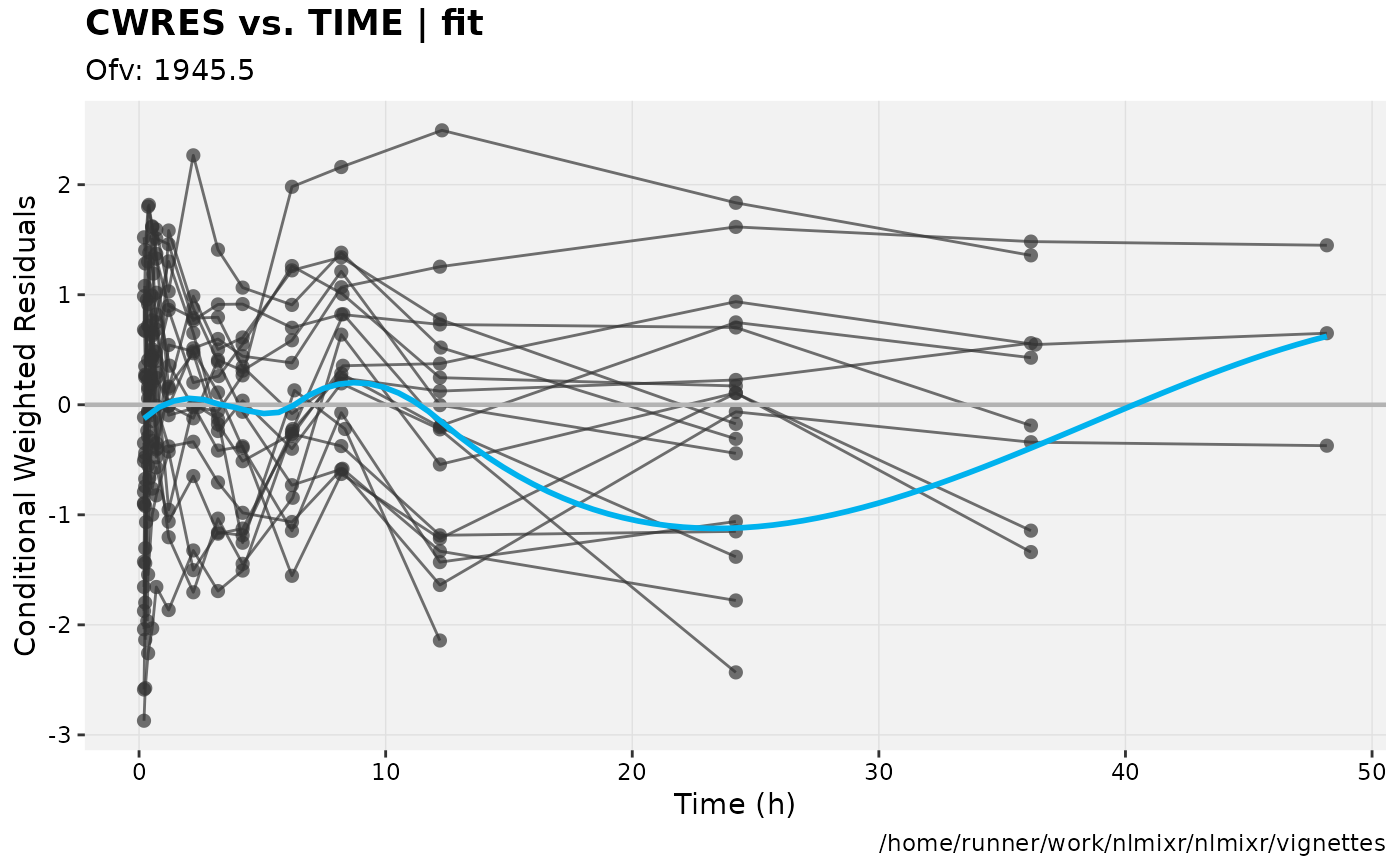
print(res_vs_idv(xpdb) +
ylab("Conditional Weighted Residuals") +
xlab("Time (h)"));
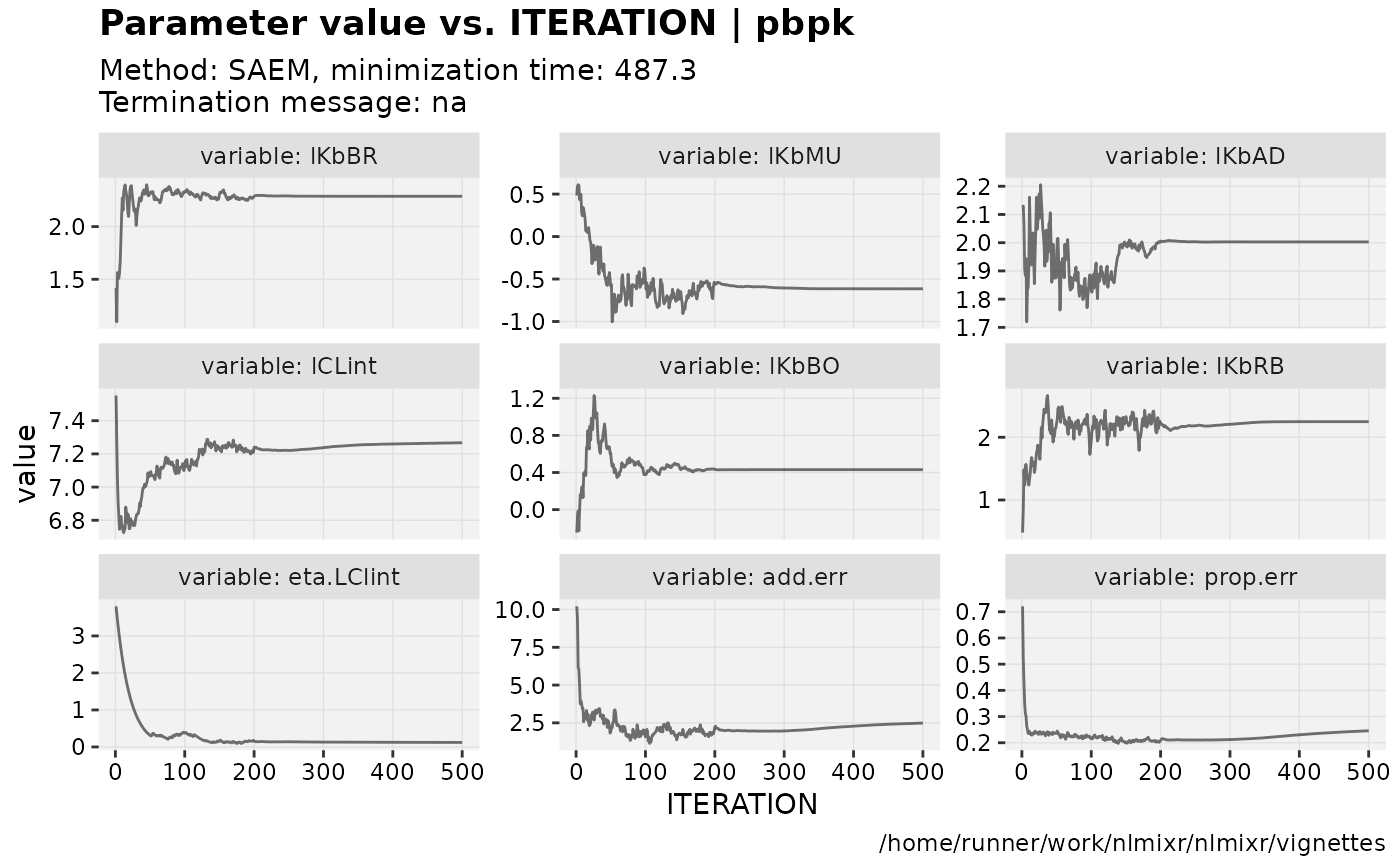
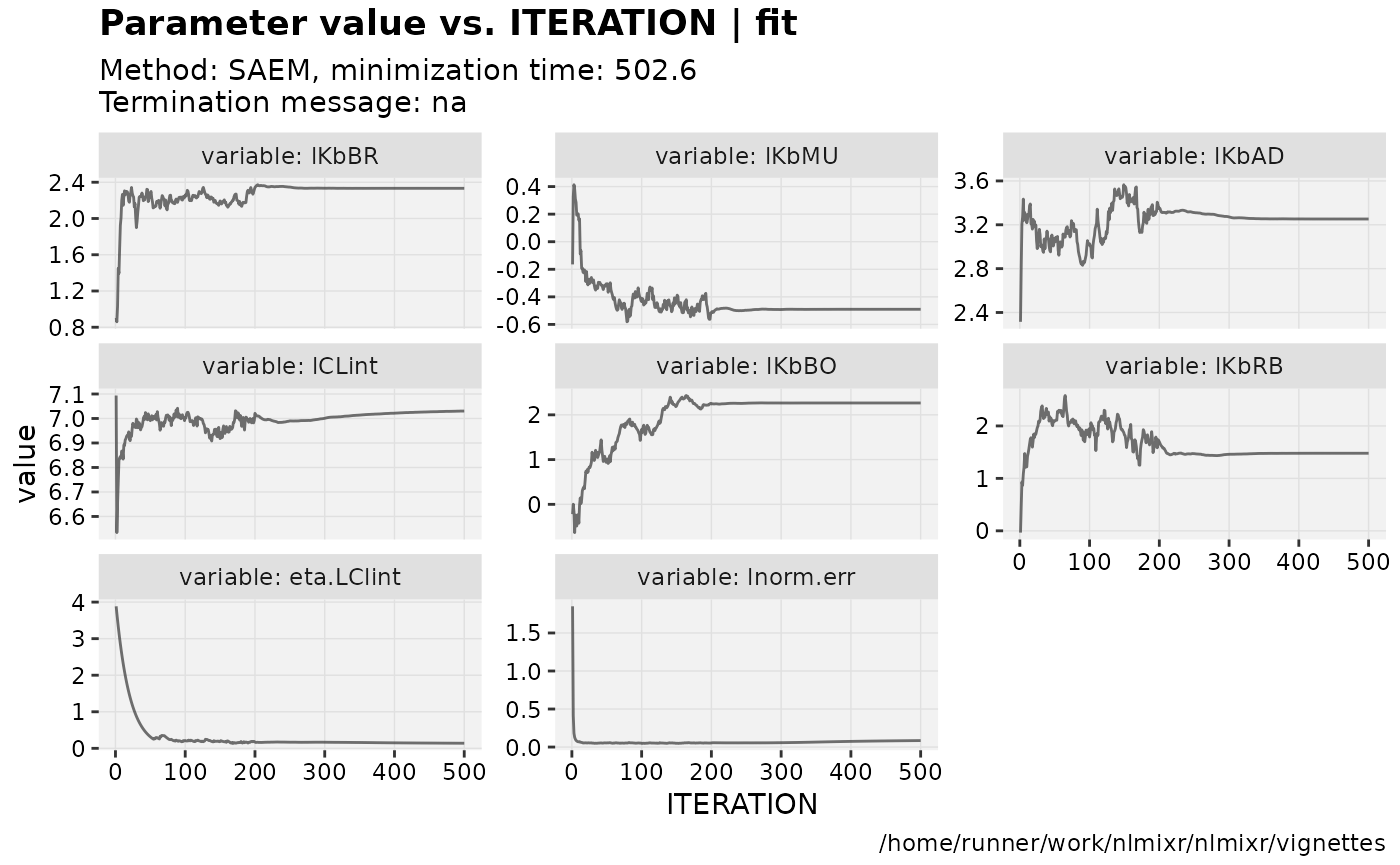
if (!is.null(fit$saem)){
print(prm_vs_iteration(xpdb));
}
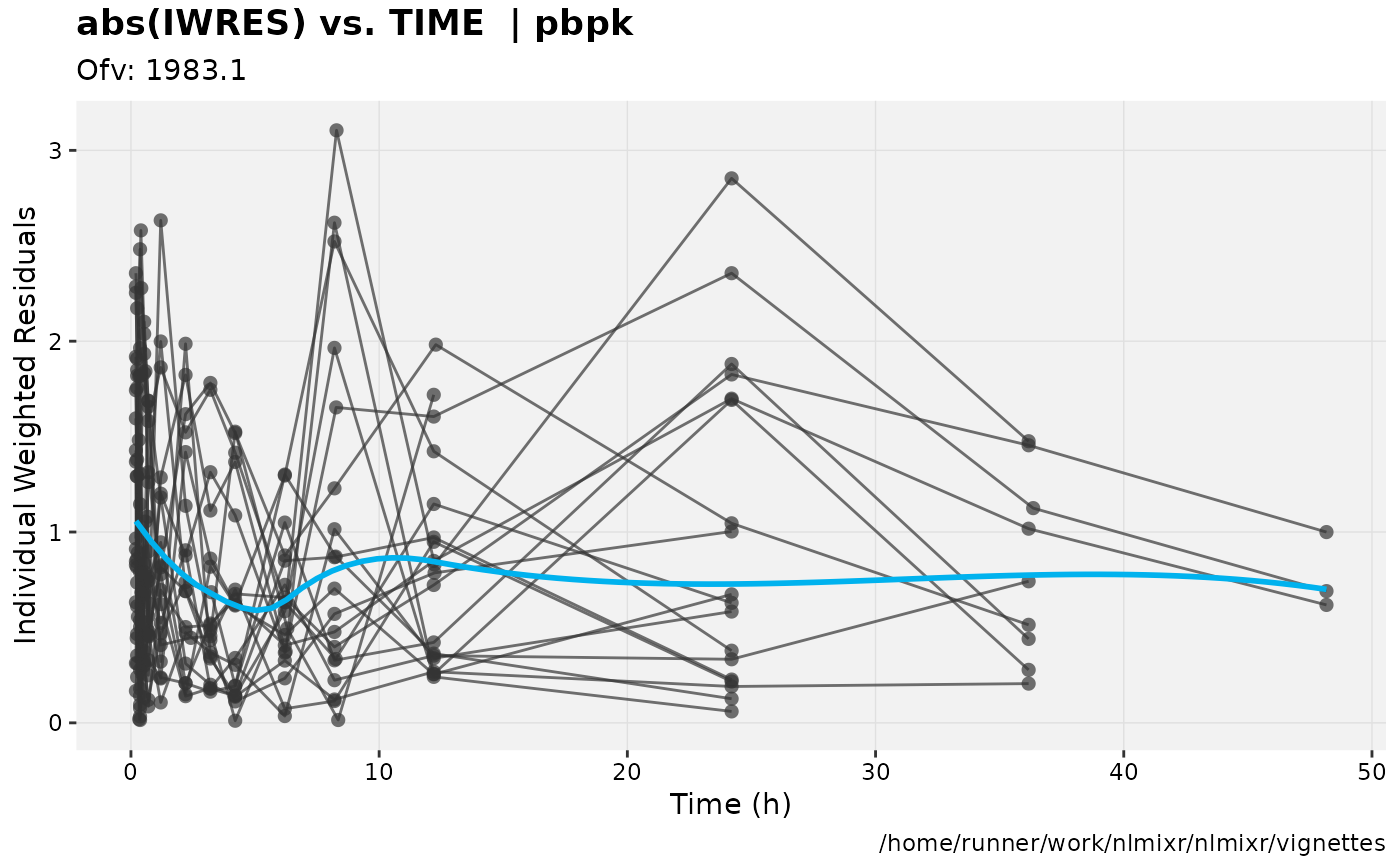
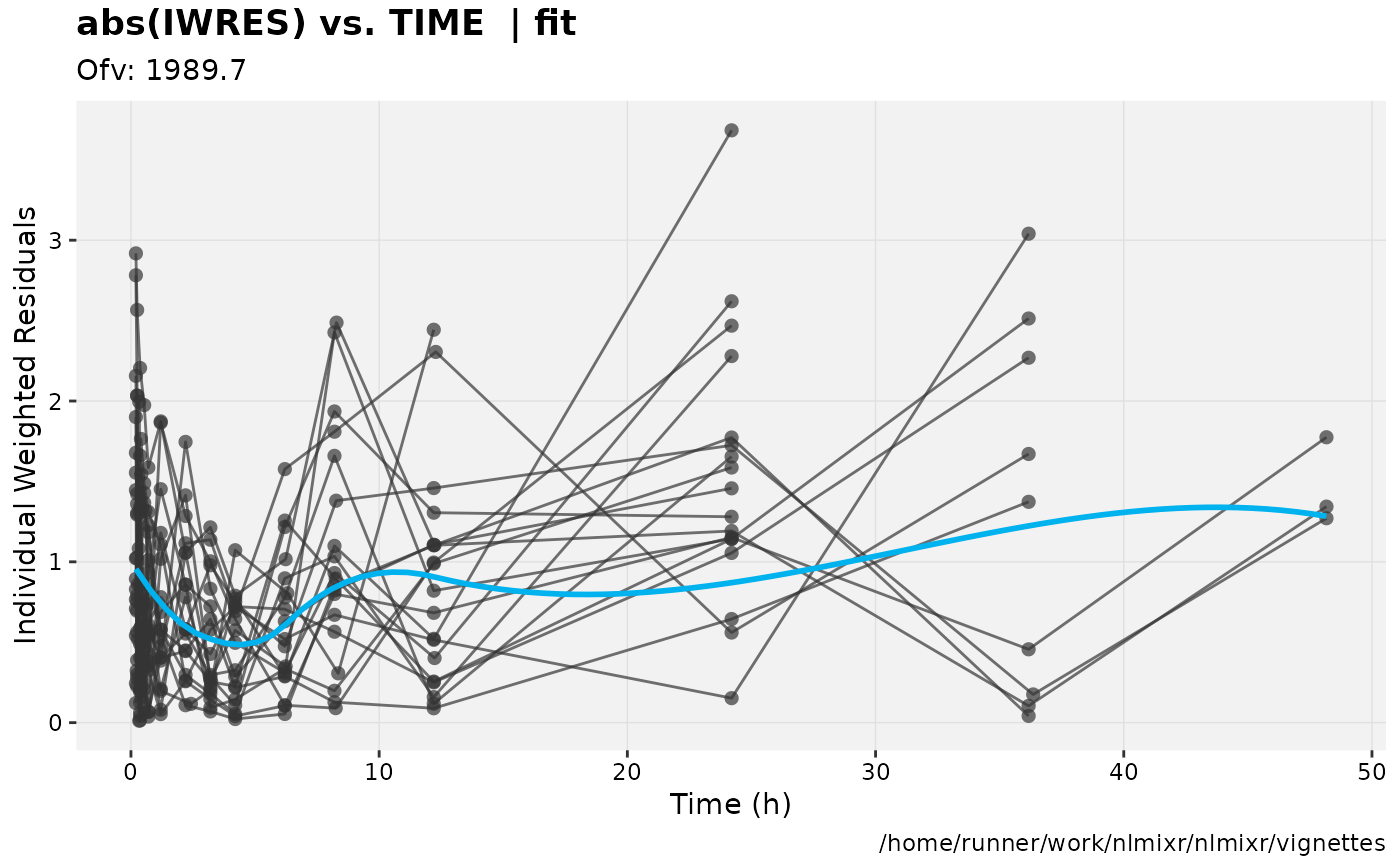
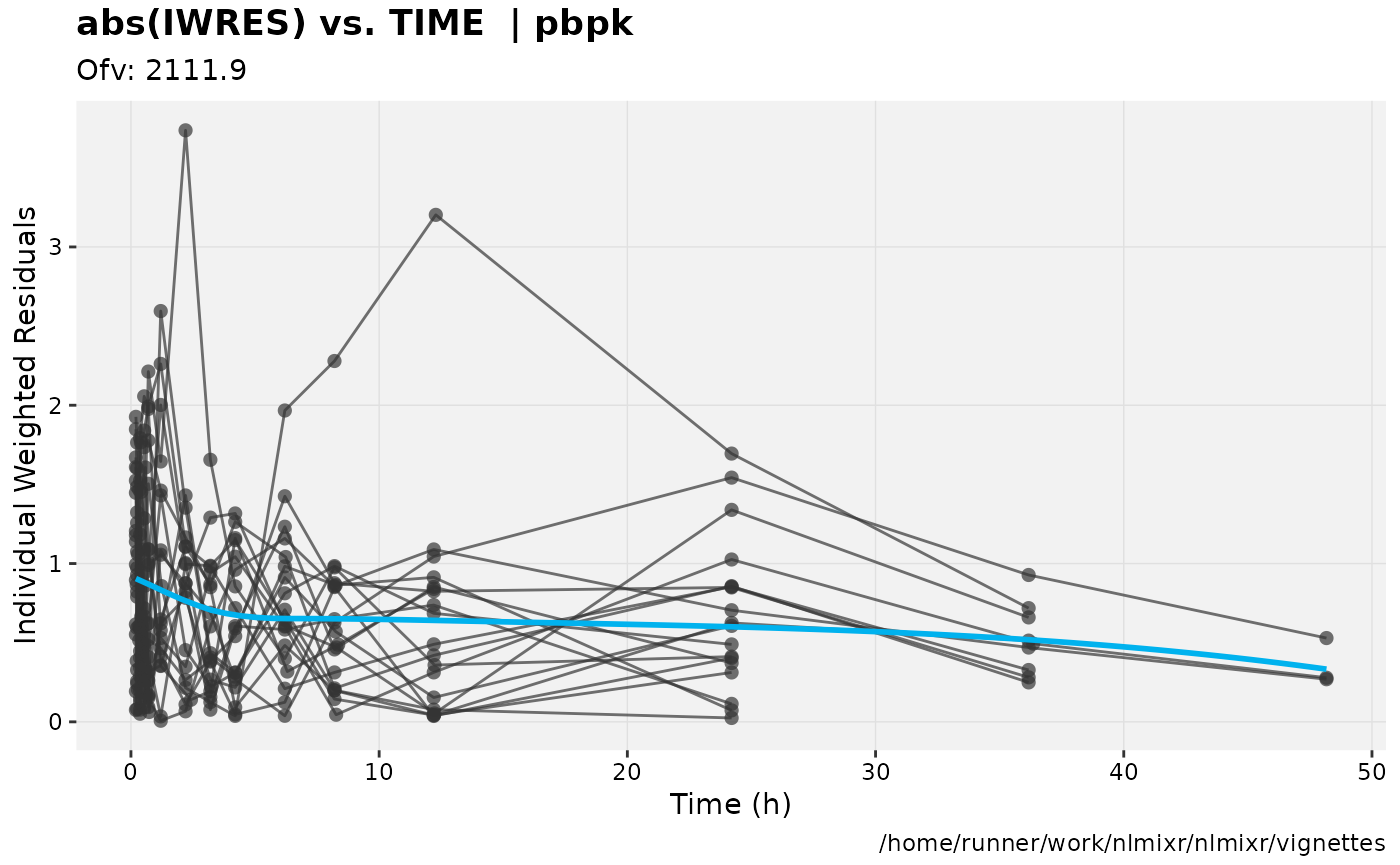
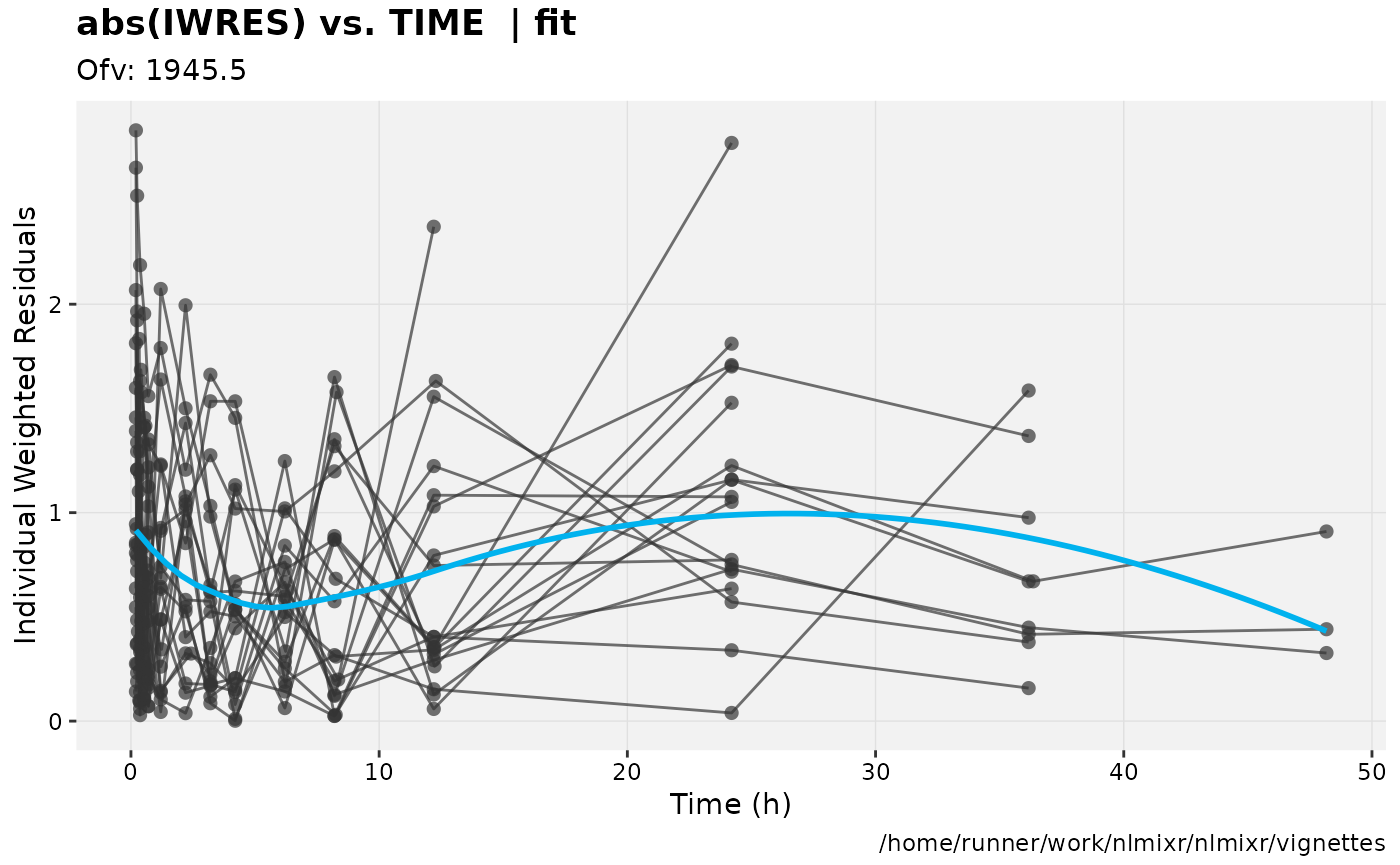
print(absval_res_vs_idv(xpdb, res = 'IWRES') +
ylab("Individual Weighted Residuals") +
xlab("Time (h)"))
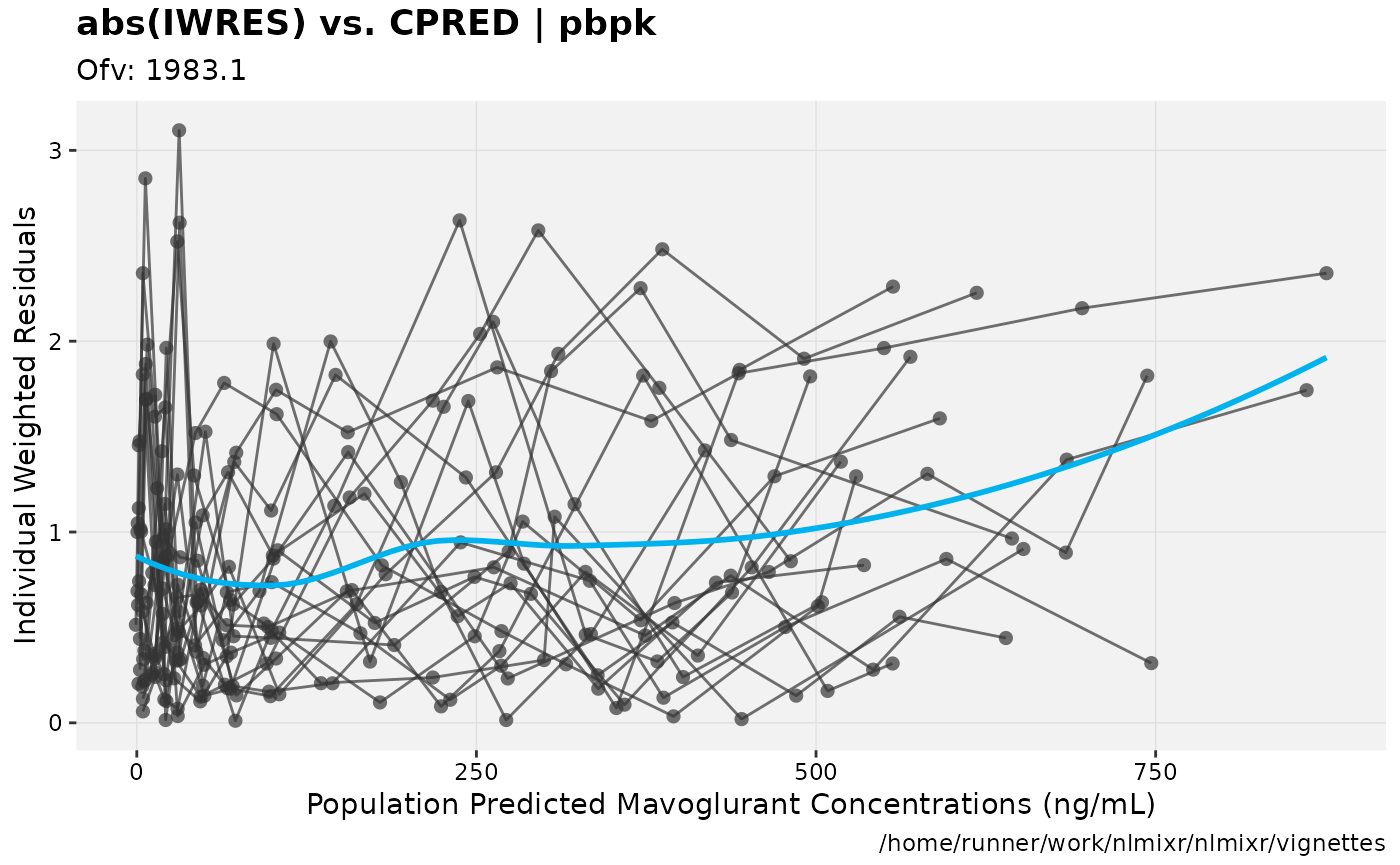
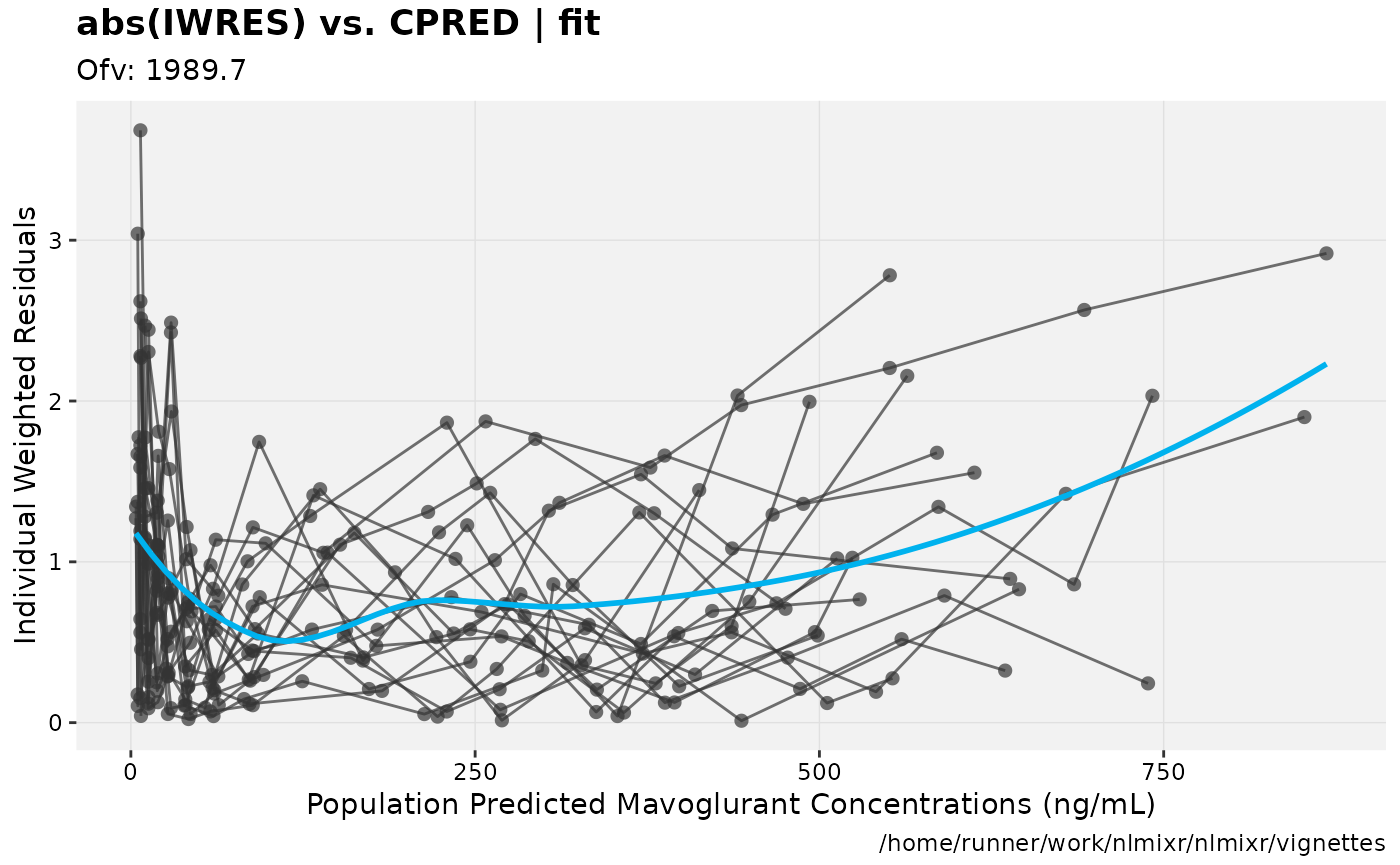
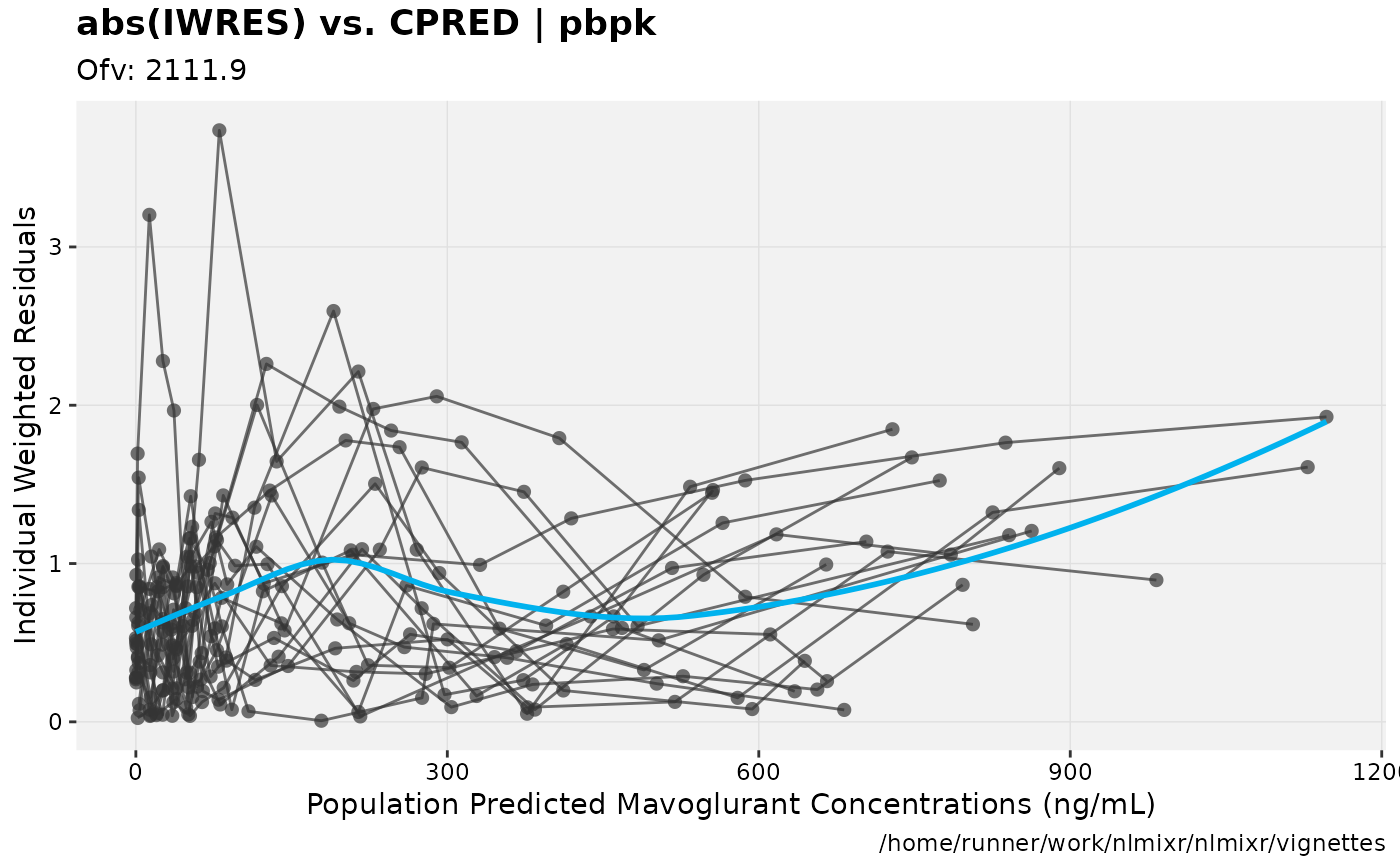
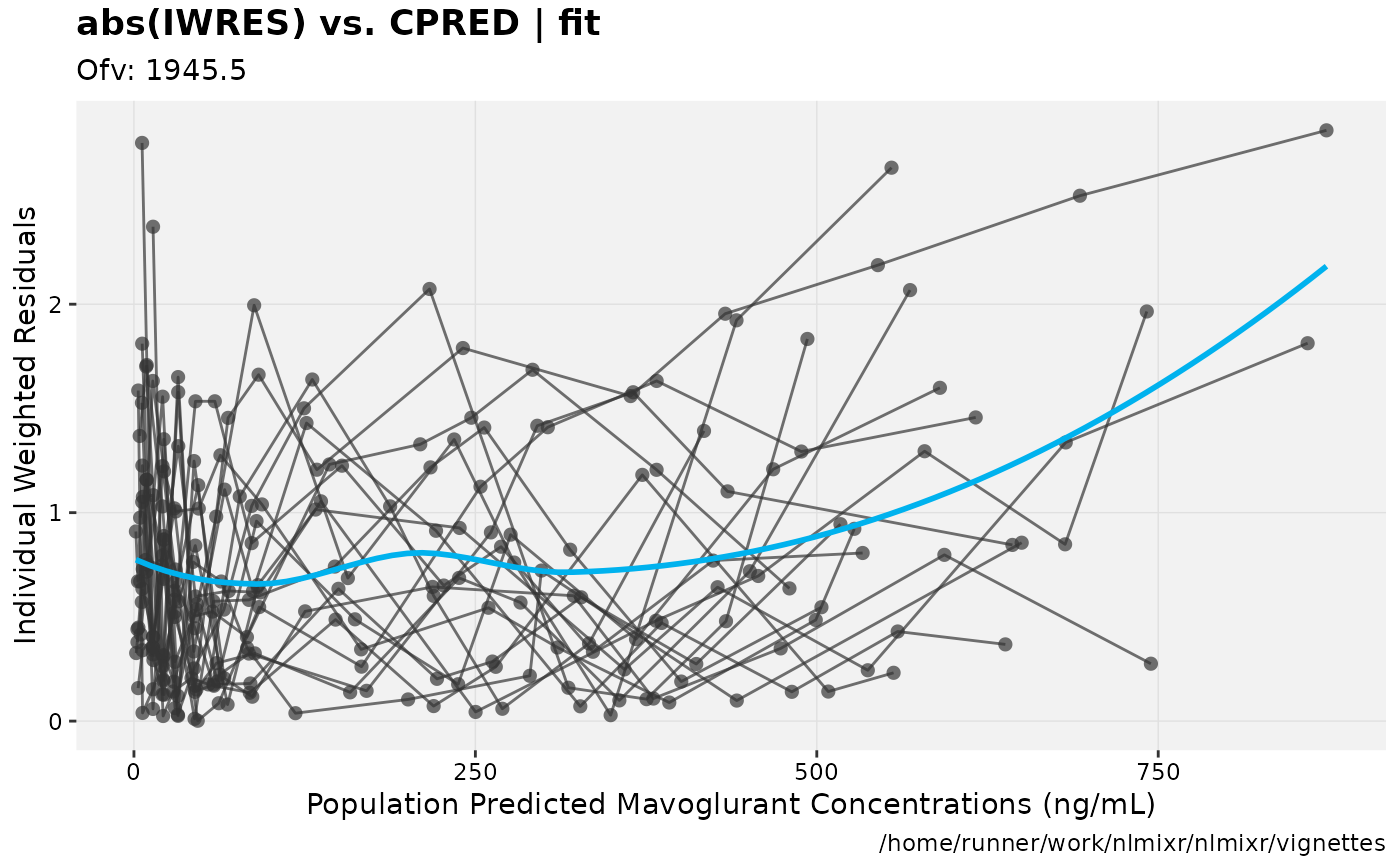
print(absval_res_vs_pred(xpdb, res = 'IWRES') +
ylab("Individual Weighted Residuals") +
xlab("Population Predicted Mavoglurant Concentrations (ng/mL)"))
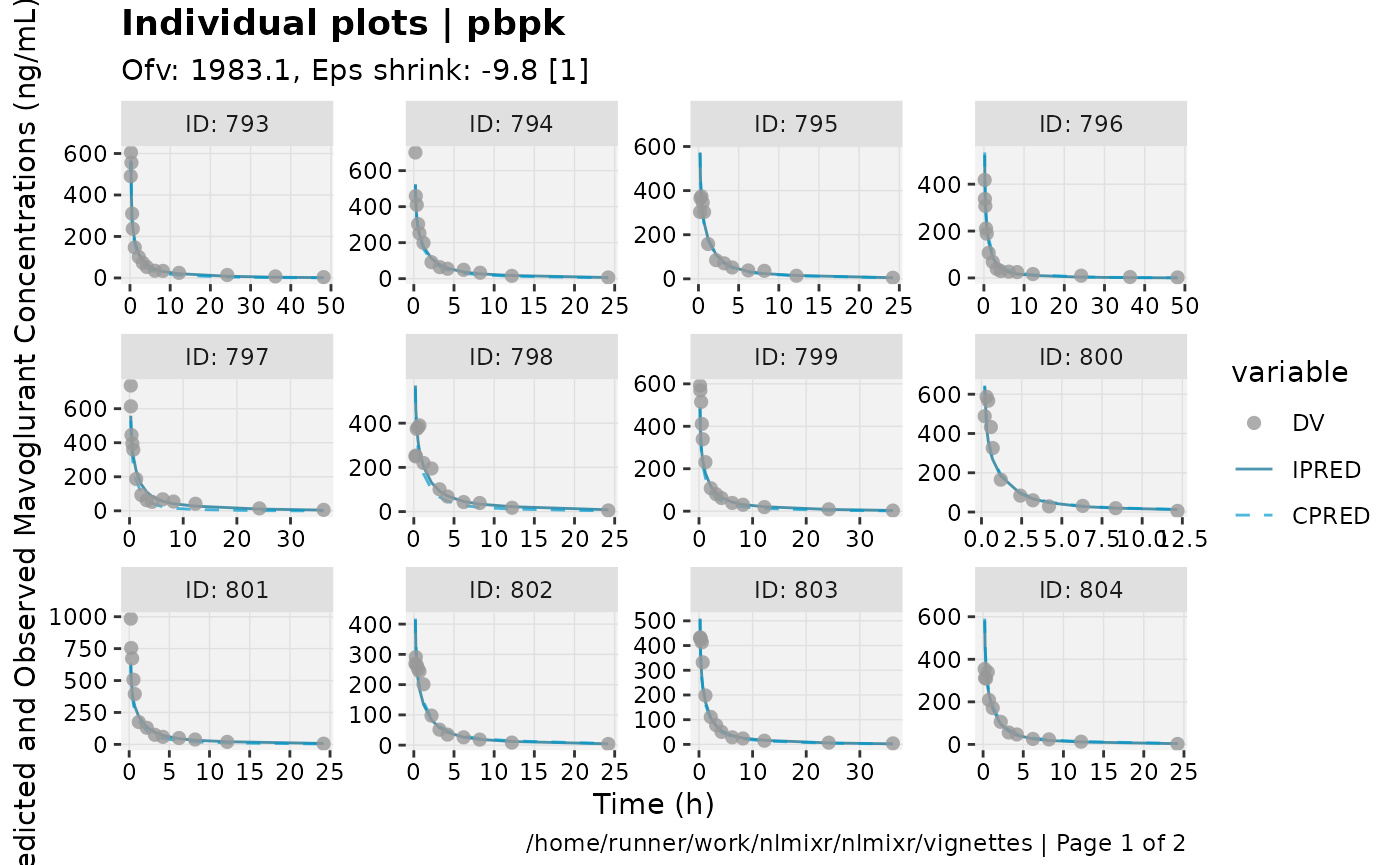
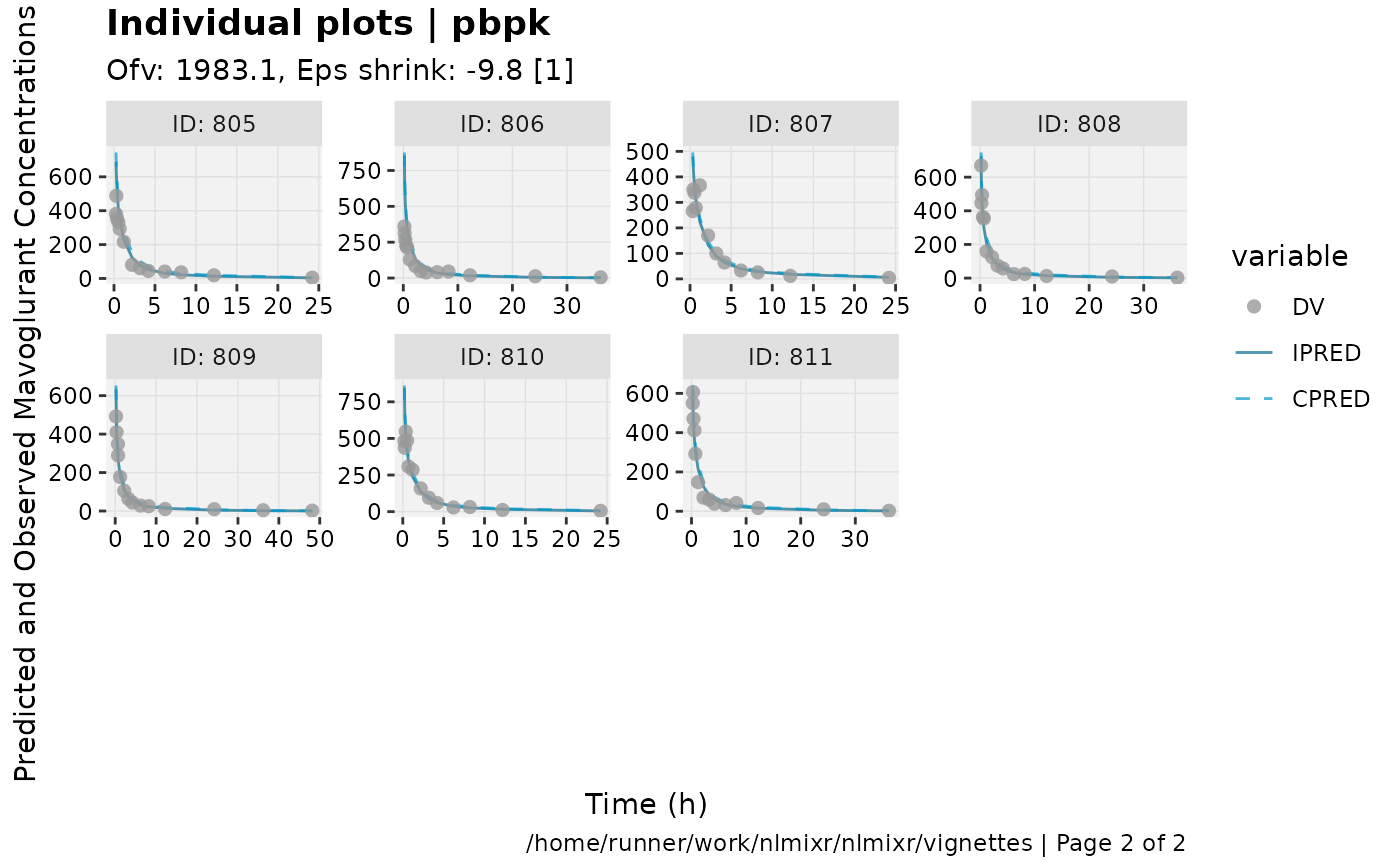
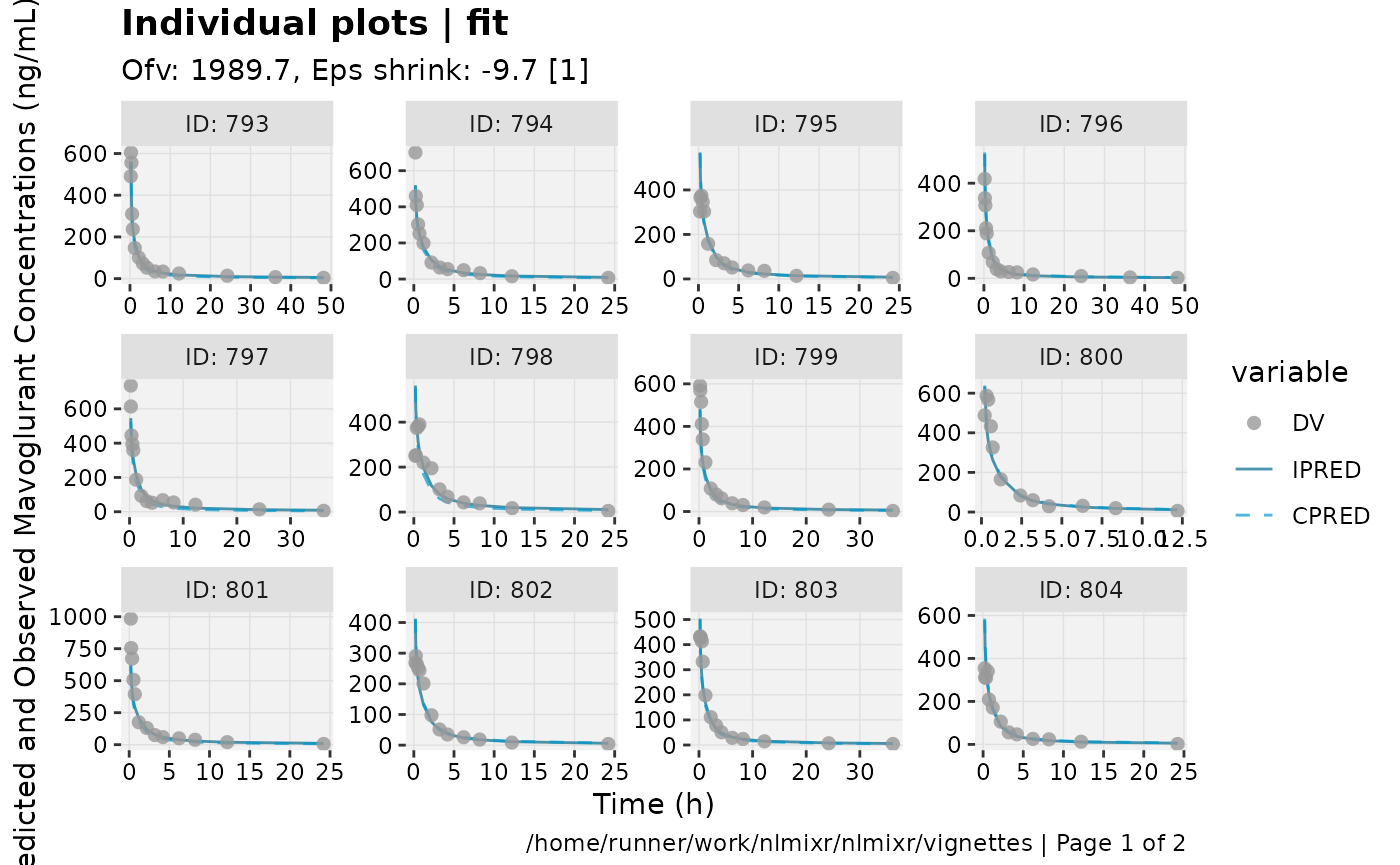
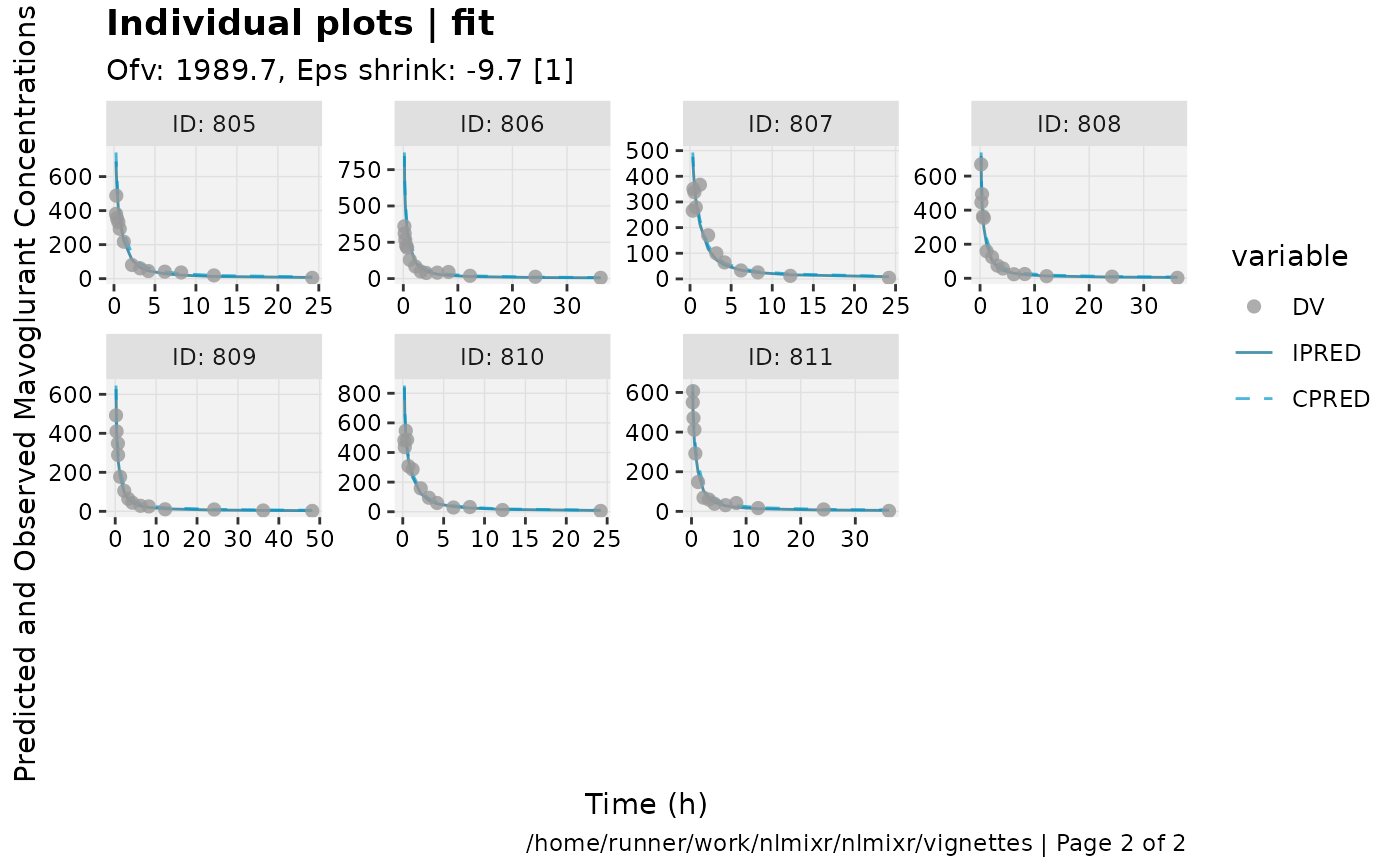
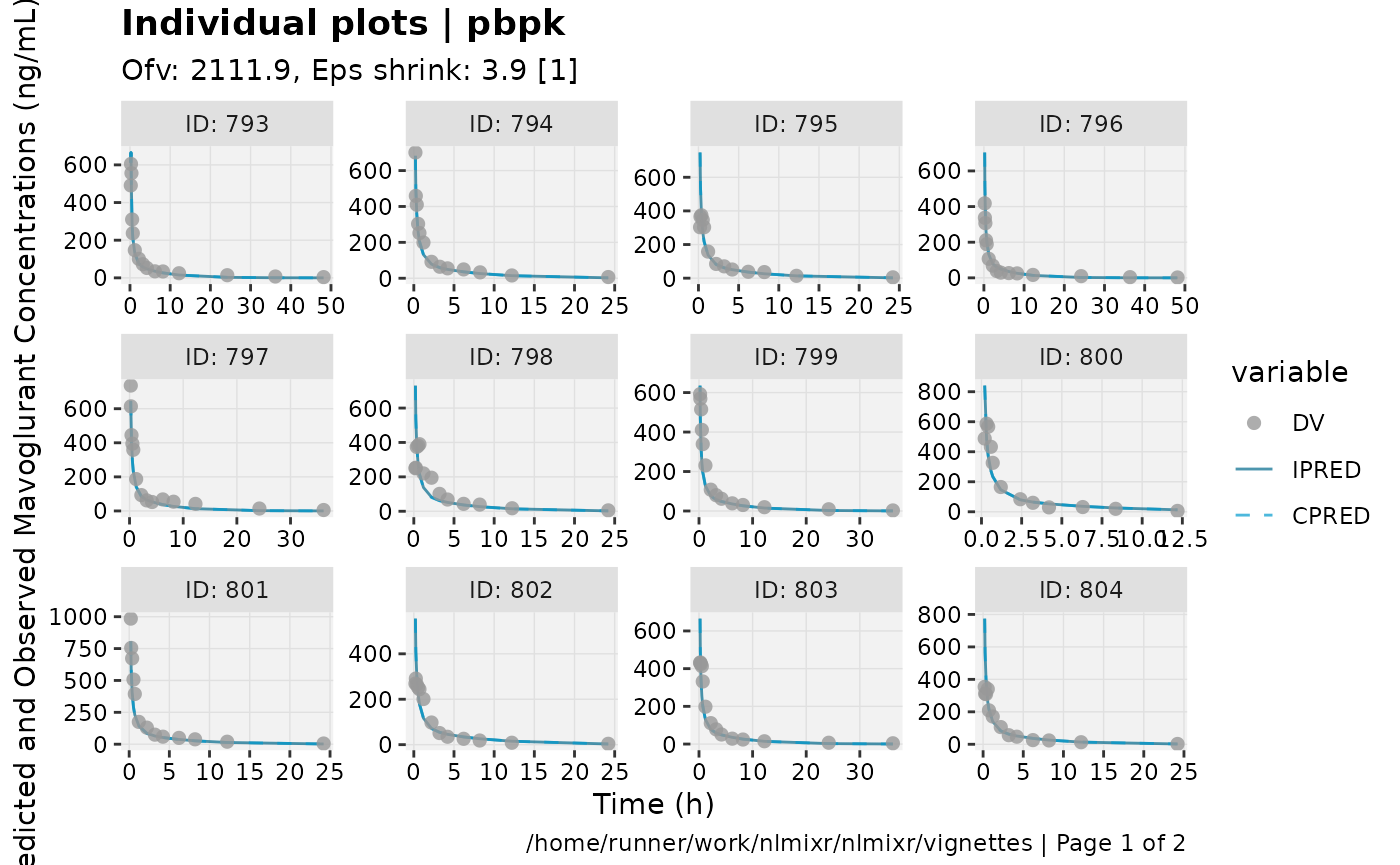
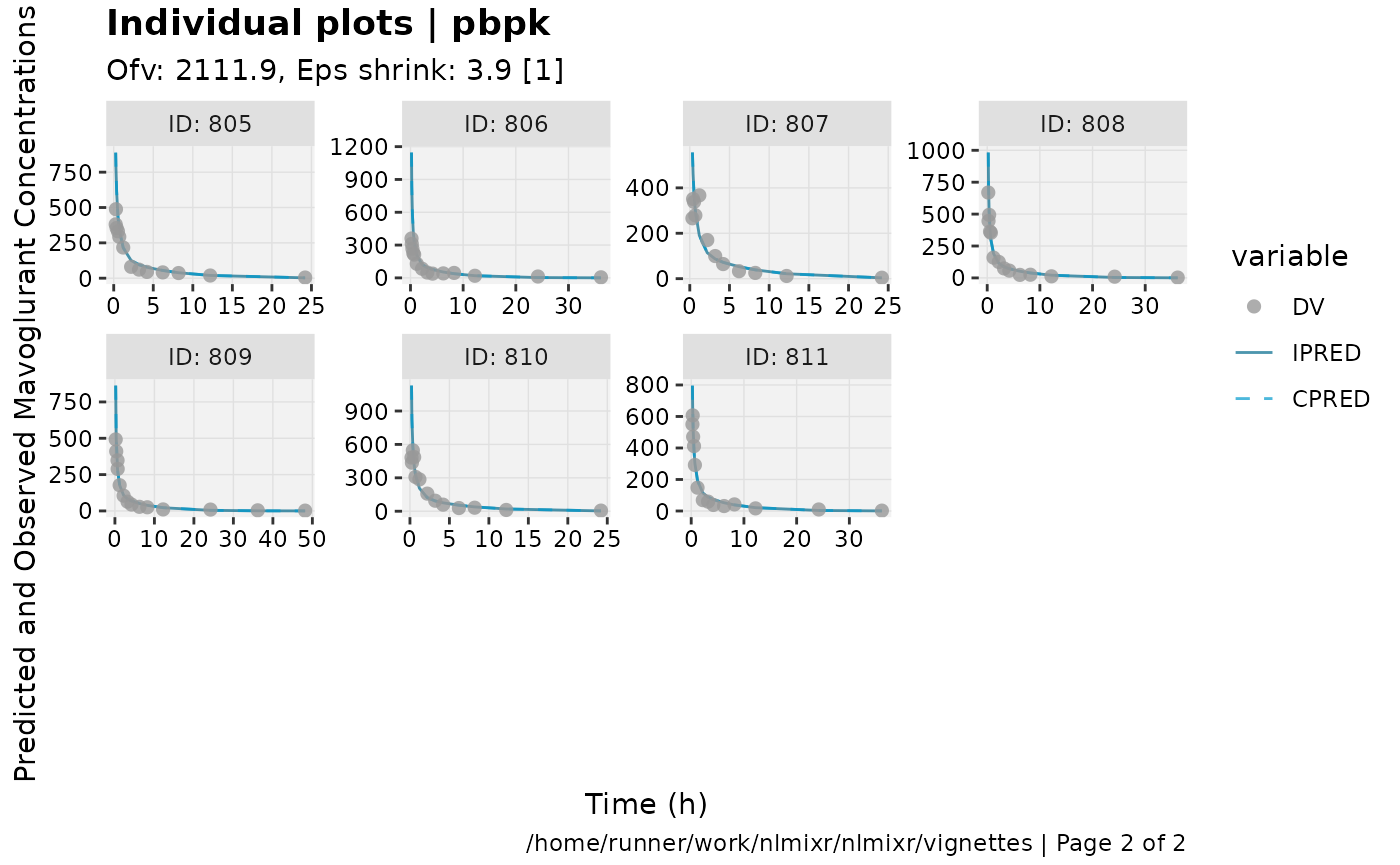
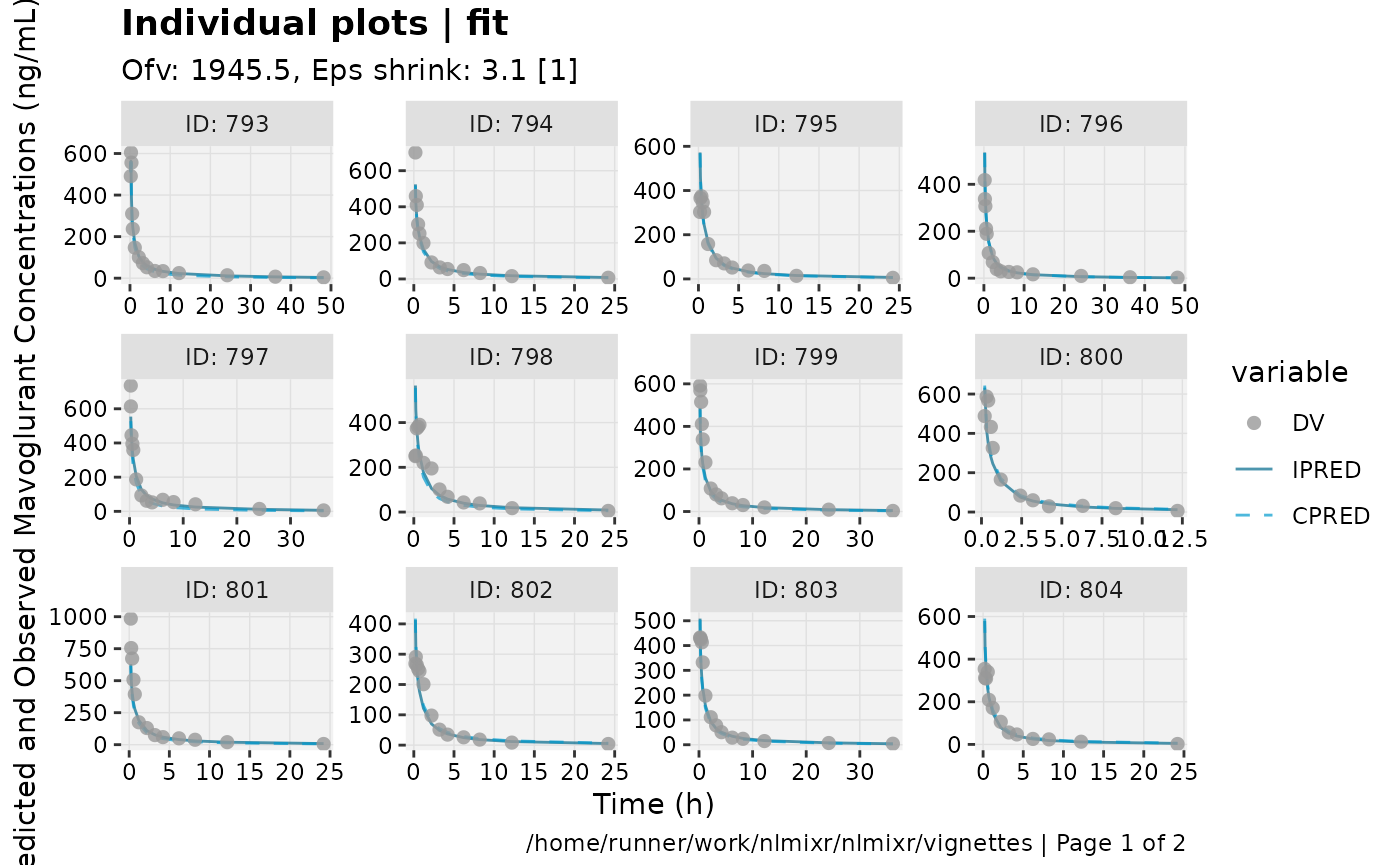
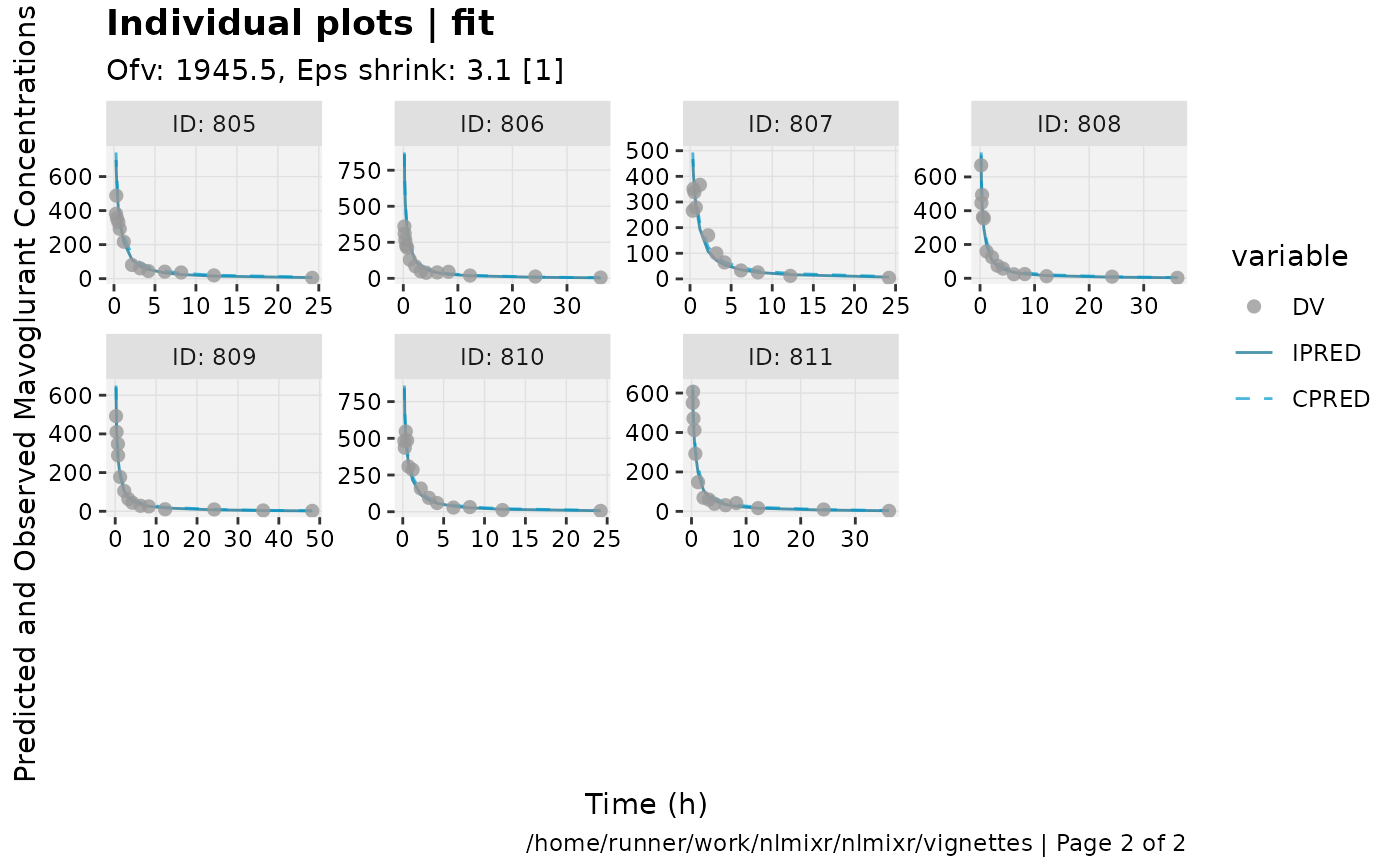
print(ind_plots(xpdb, nrow=3, ncol=4) +
ylab("Predicted and Observed Mavoglurant Concentrations (ng/mL)") +
xlab("Time (h)"))
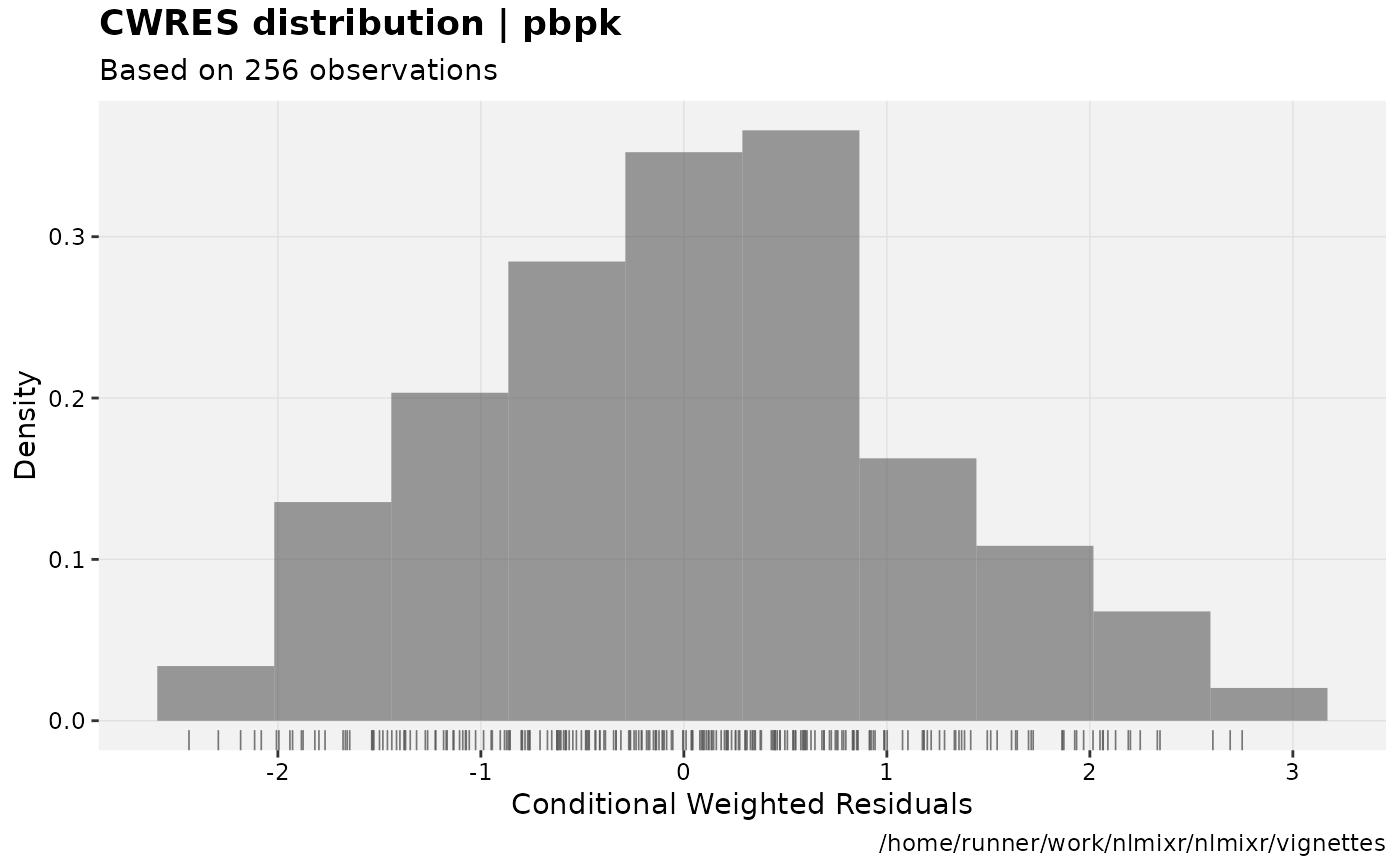
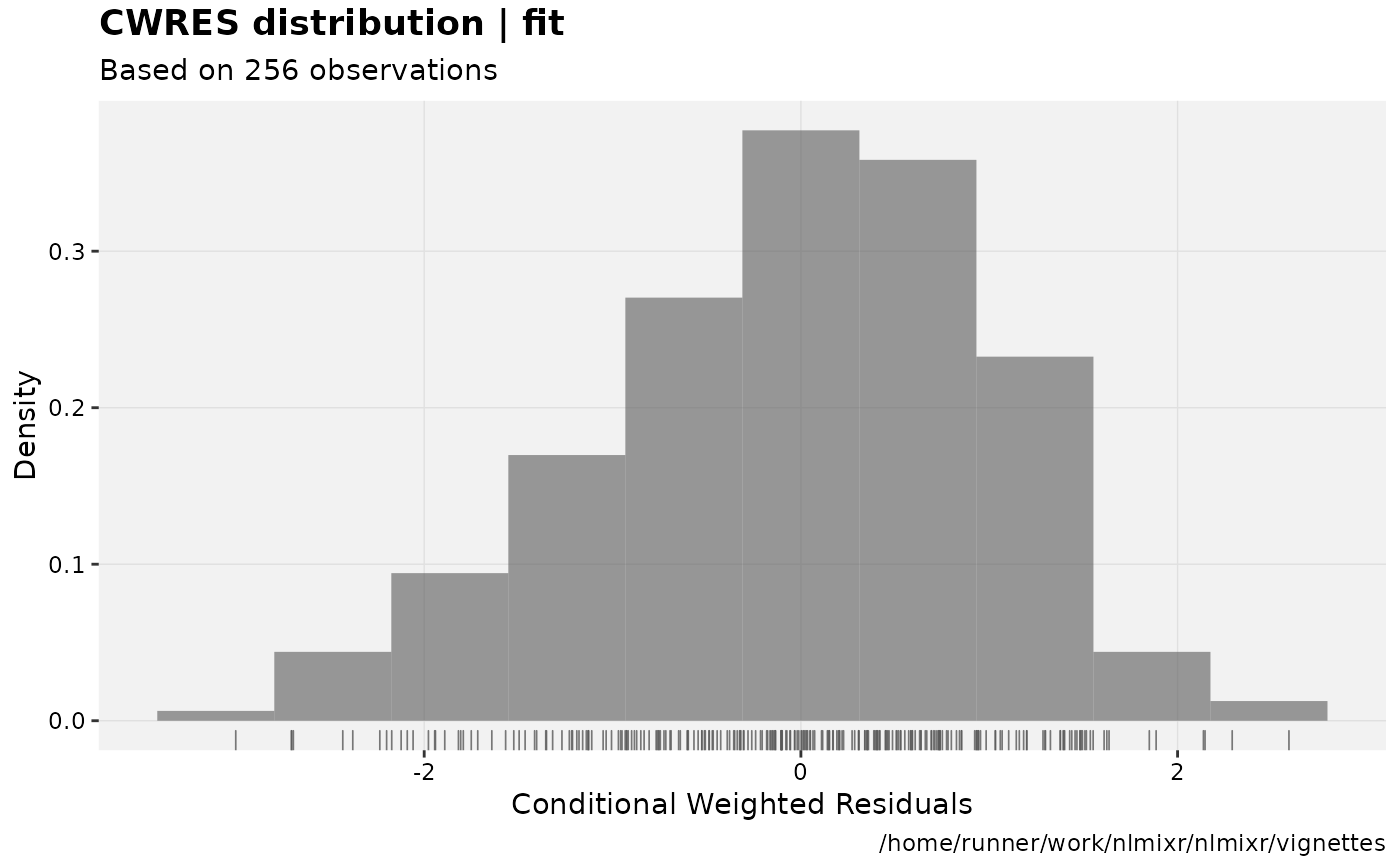
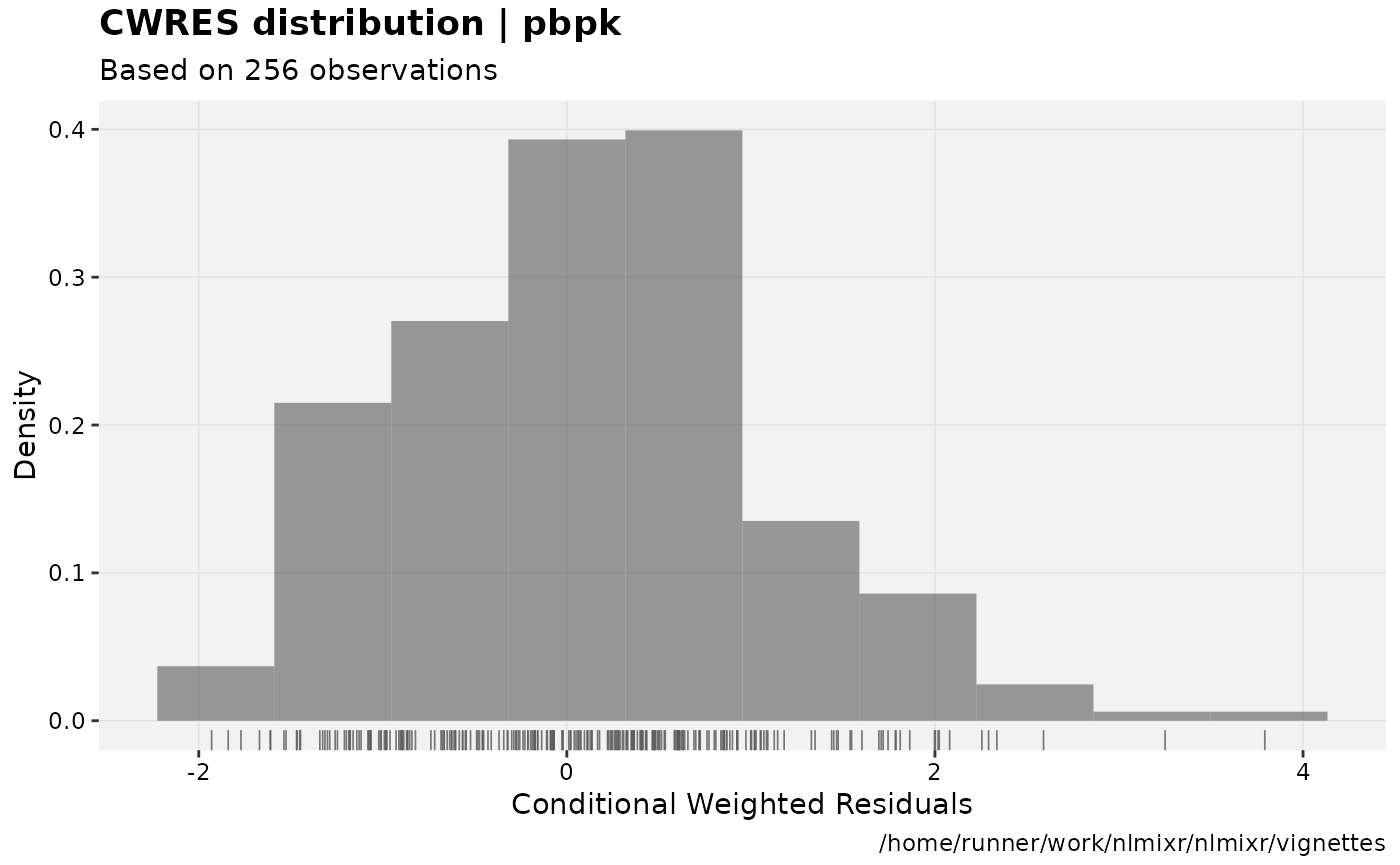
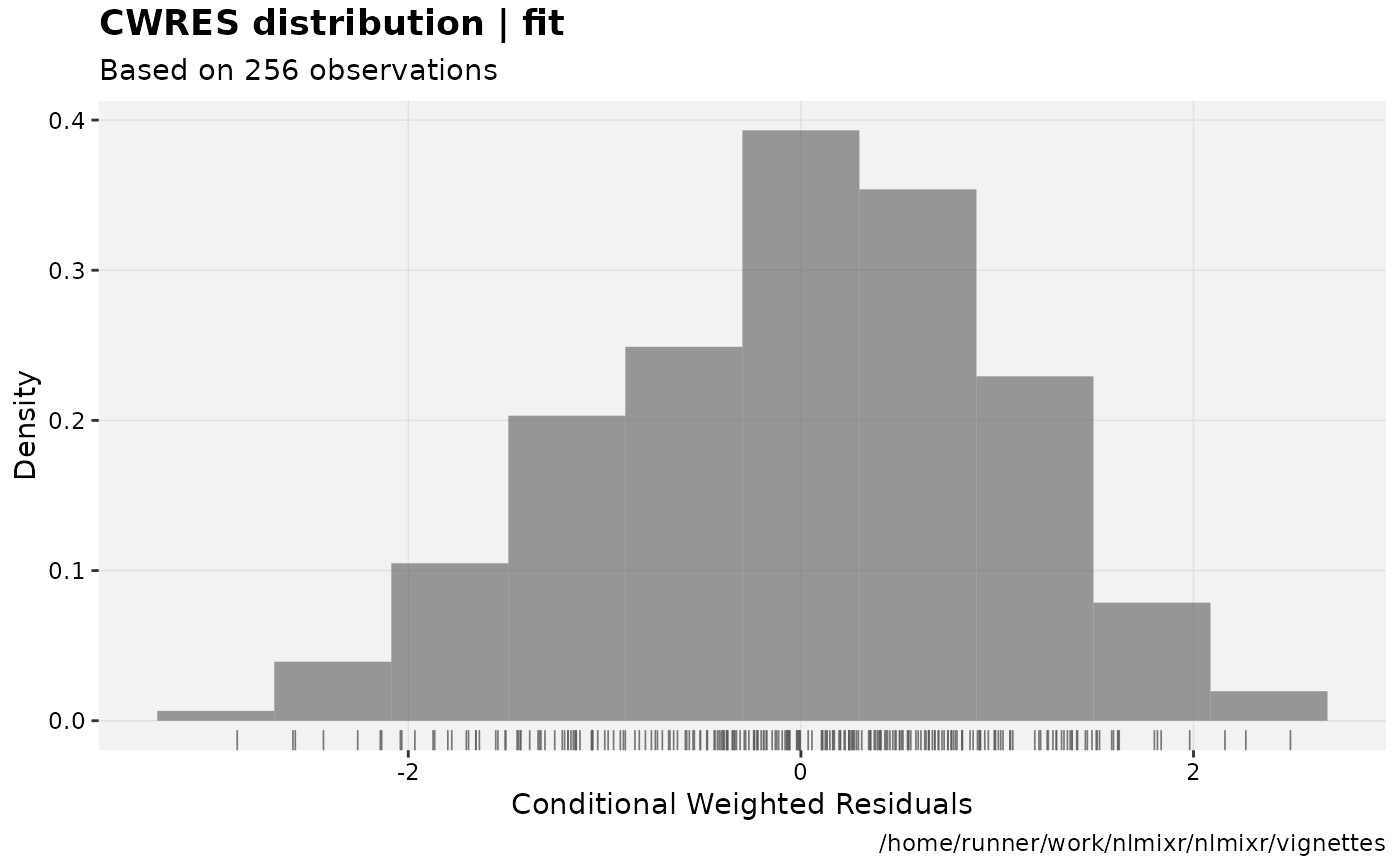
print(res_distrib(xpdb) +
ylab("Density") +
xlab("Conditional Weighted Residuals"));
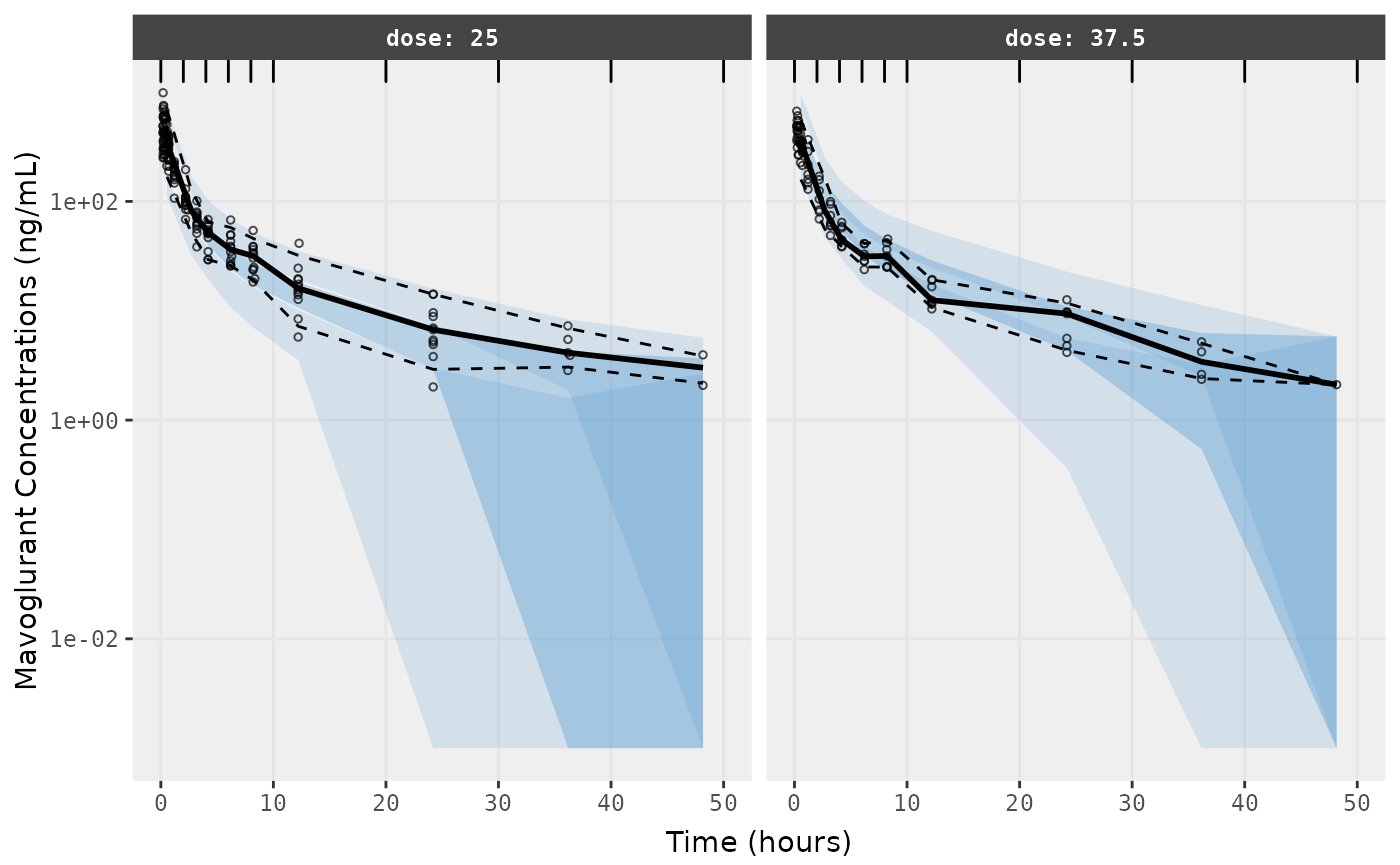
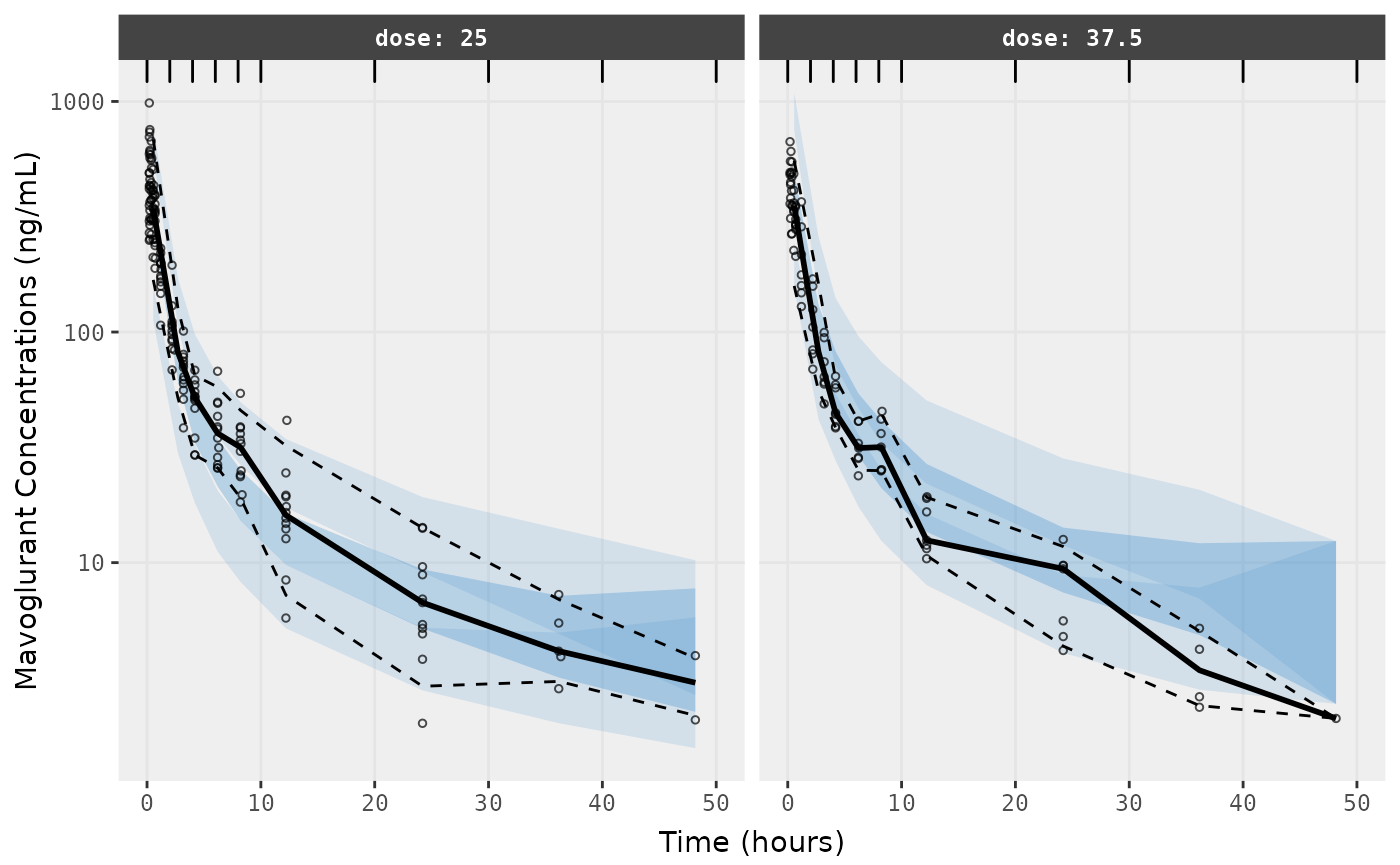
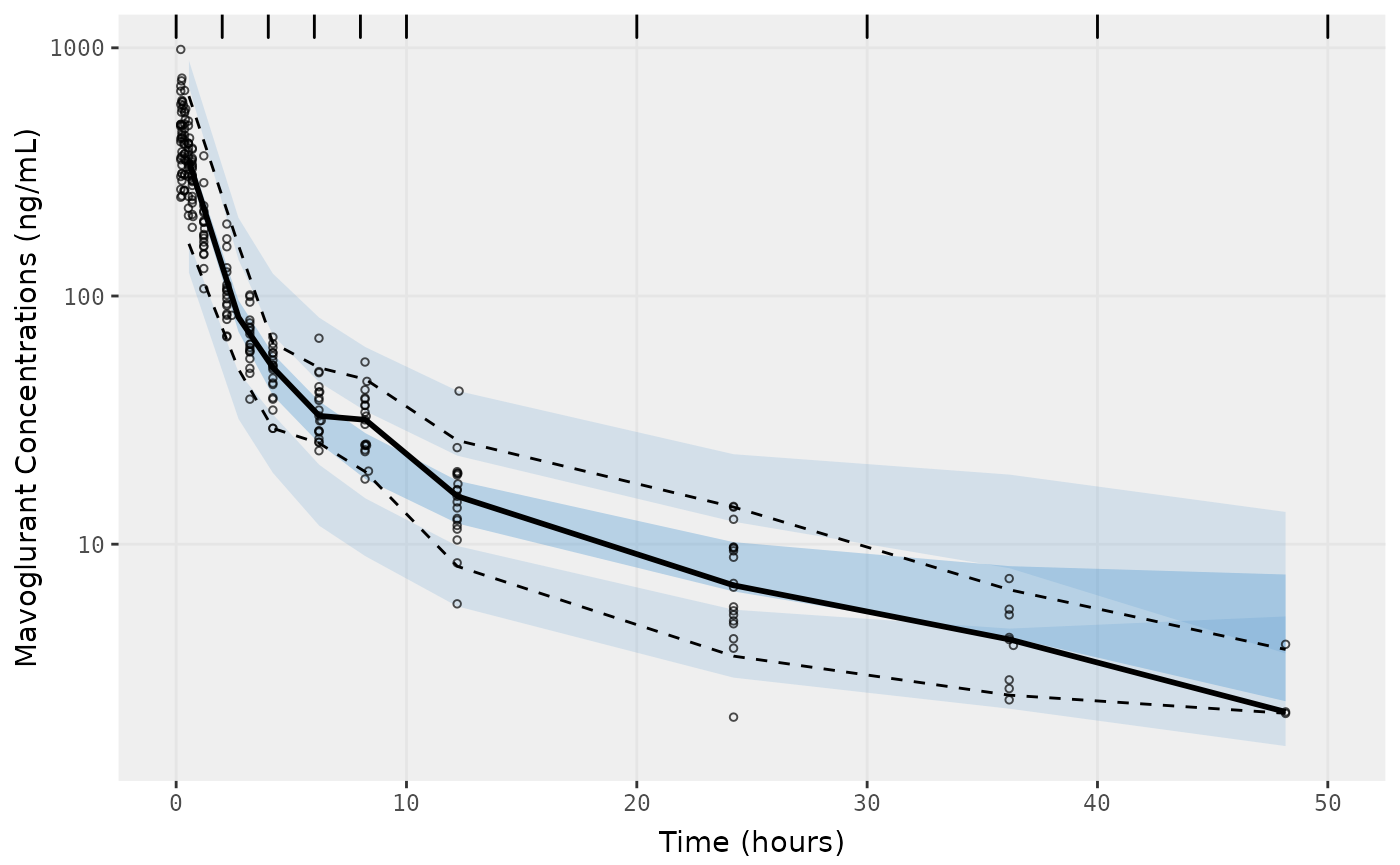
##Visual Predictive Checks
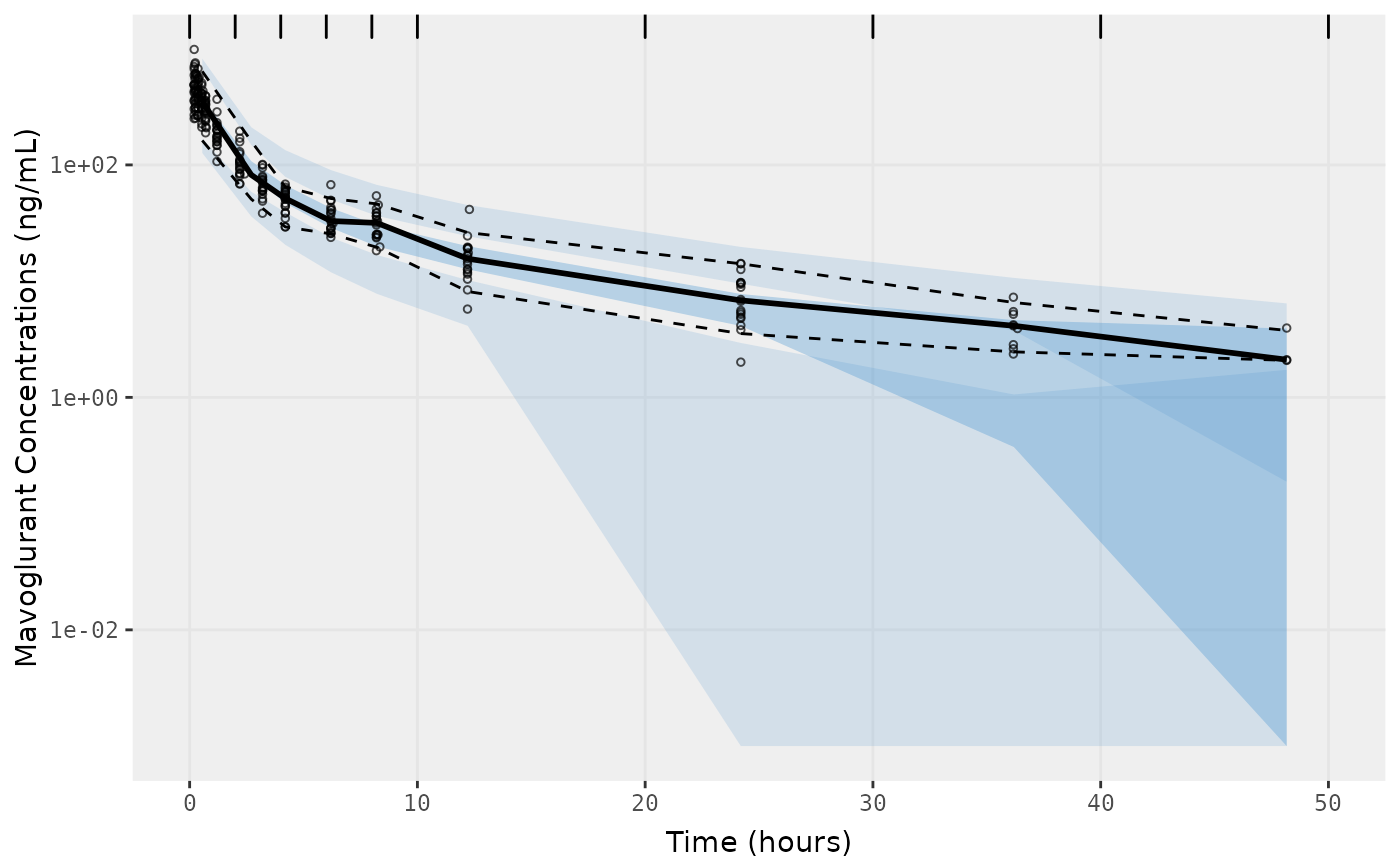
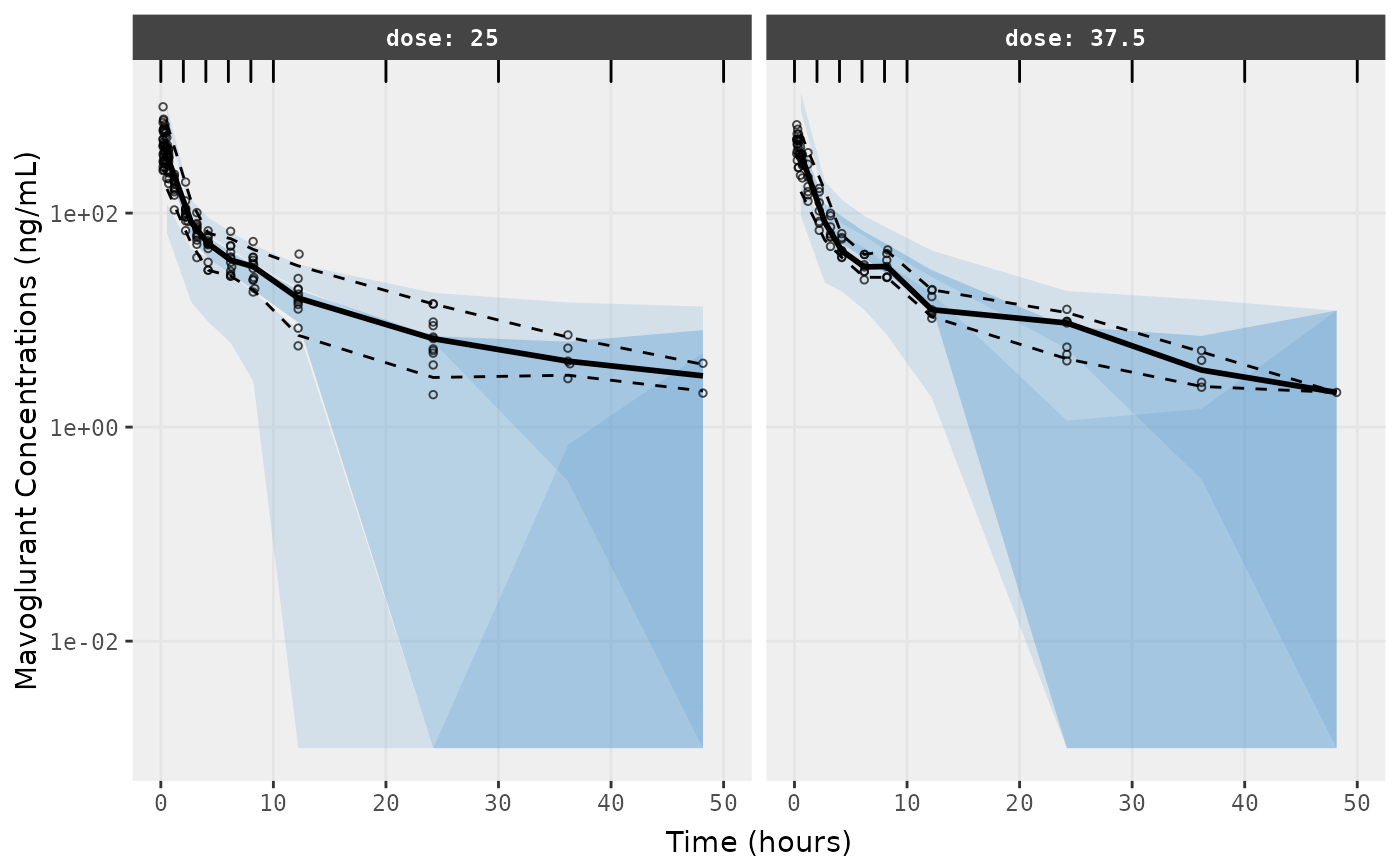
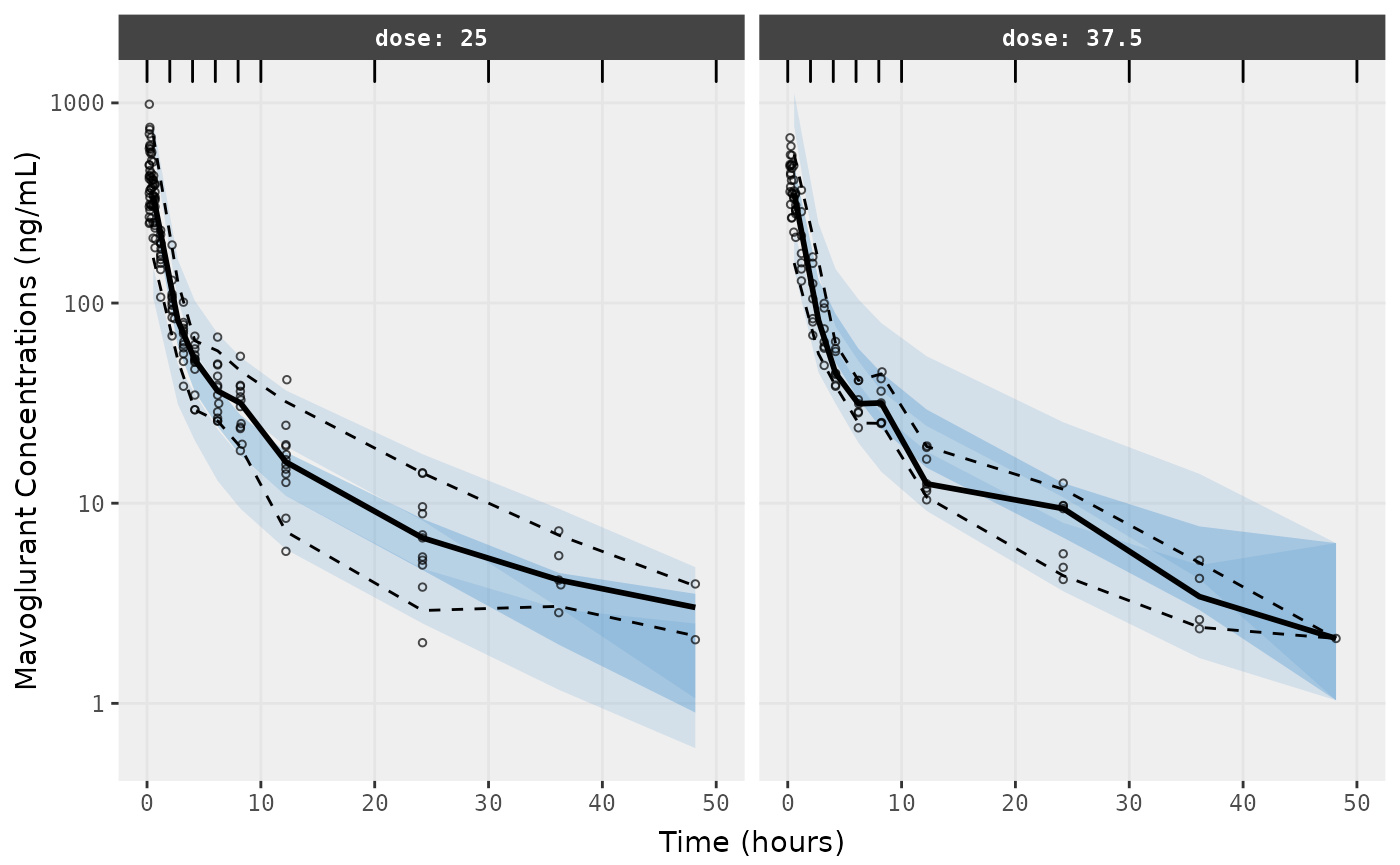
f1 <- vpc.ui(fit,n=500,stratify=c("dose"), show=list(obs_dv=T), log_y=TRUE,
bins = c(0, 2, 4, 6, 8, 10, 20, 30, 40, 50),
ylab = "Mavoglurant Concentrations (ng/mL)",
xlab = "Time (hours)")
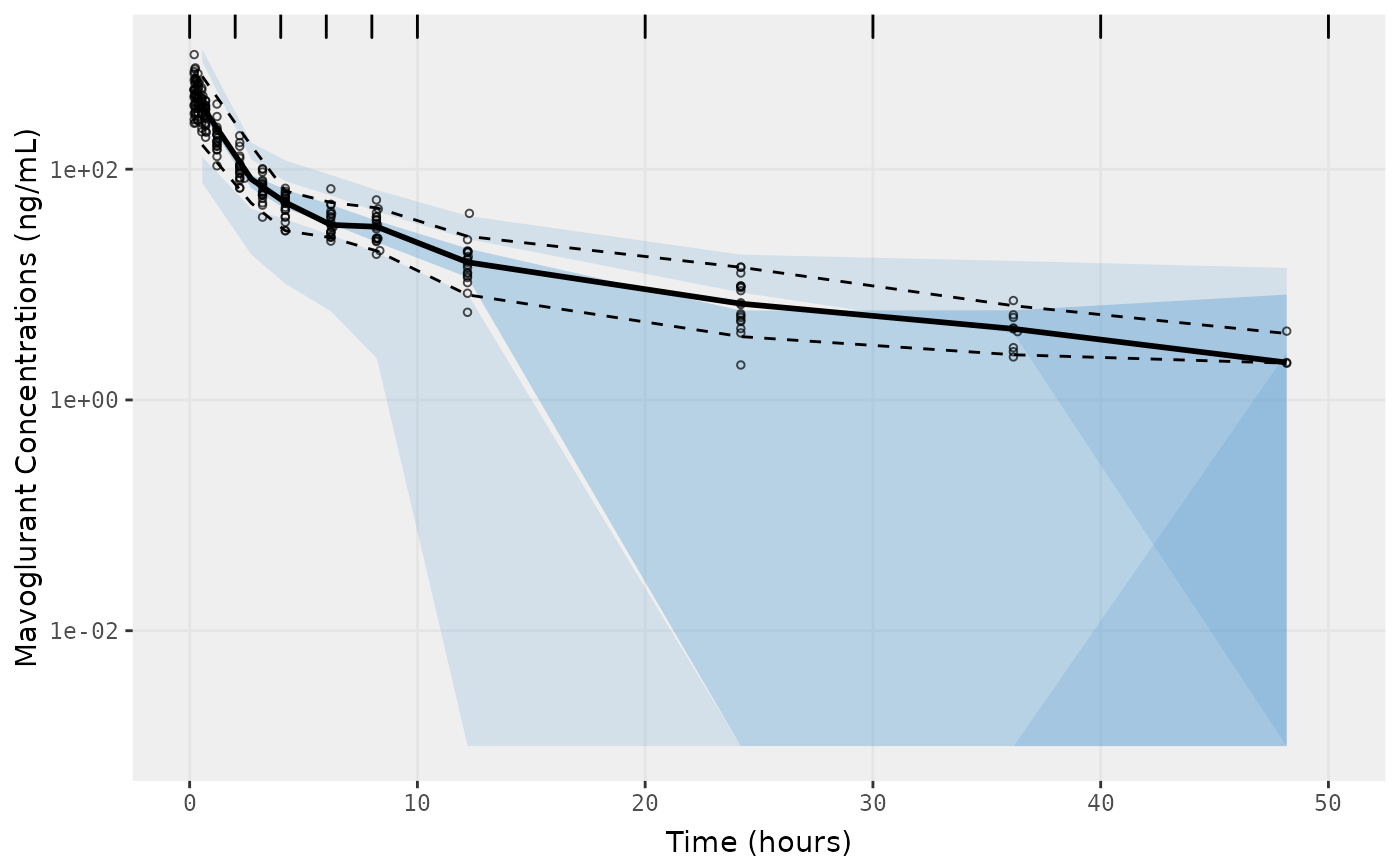
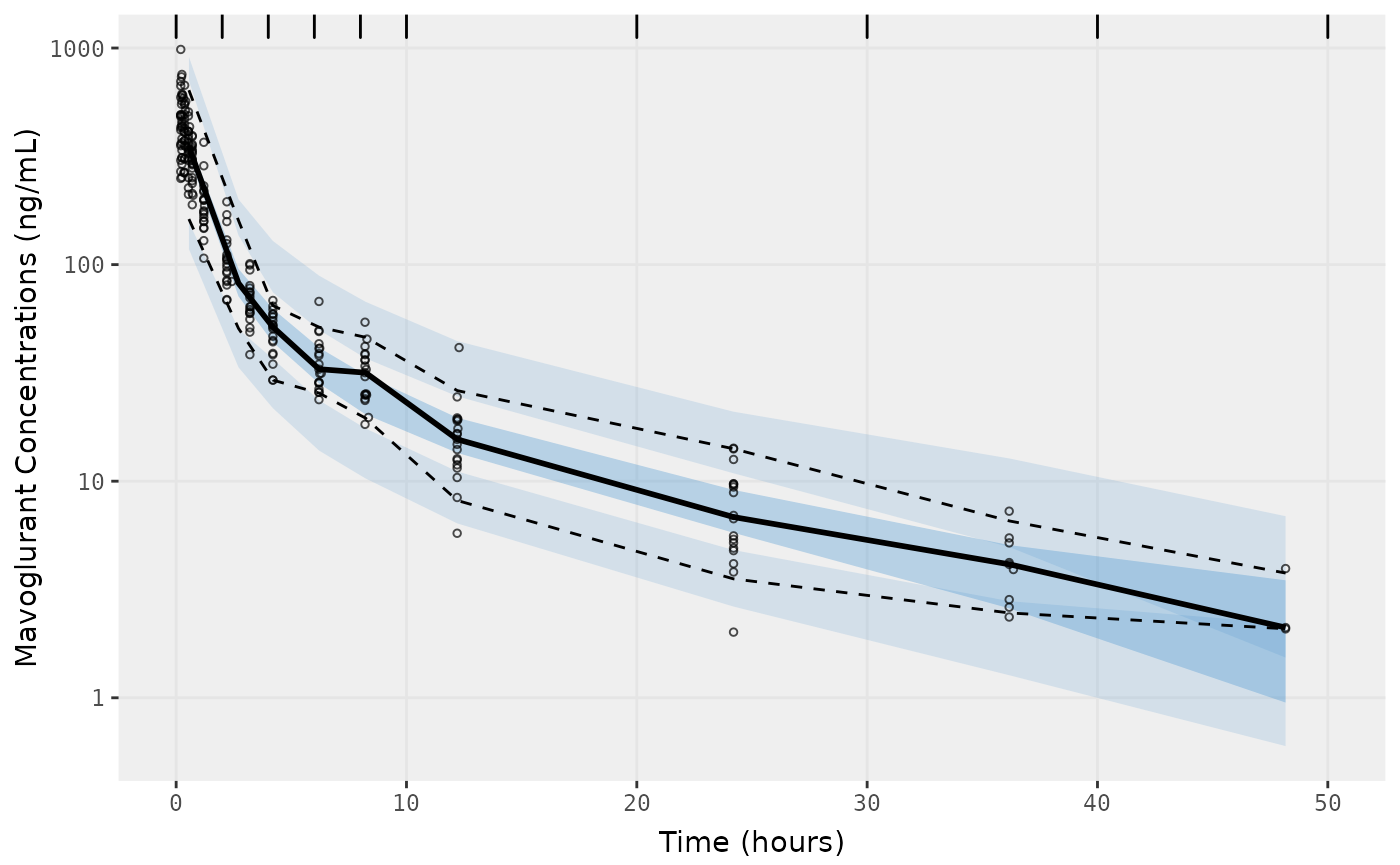
f2 <- vpc.ui(fit,n=500, show=list(obs_dv=T), bins = c(0, 2, 4, 6, 8, 10, 20, 30, 40, 50), log_y=TRUE,
ylab = "Mavoglurant Concentrations (ng/mL)", xlab = "Time (hours)")
plot(f1)
plot(f2)
}SAEM
Fit add+prop SAEM
fit.addProp.S <- nlmixr(pbpk, dat, est="saem", control=list(print=0),
table=list(cwres=TRUE, npde=TRUE))
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [1] "WT"
gofs(fit.addProp.S);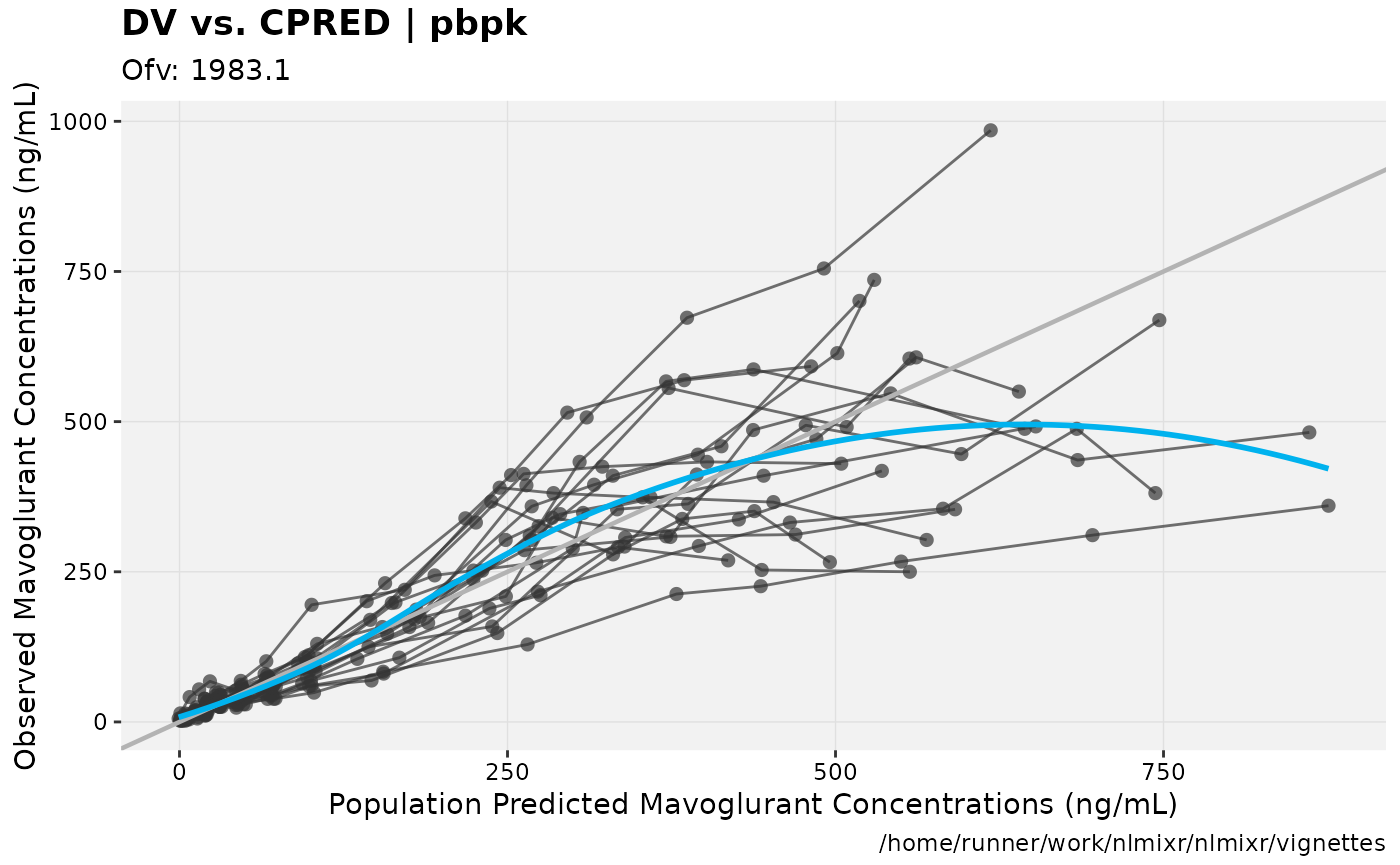
Change error to lognormal
fit.lnorm.S <- pbpk %>%
model({C15 ~ lnorm(lnorm.err)}) %>% # Change C15 to be log-normally distributed
## Requires data since piping from pbpk model
nlmixr(dat,est="saem", control=list(print=0),
table=list(cwres=TRUE, npde=TRUE))
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [1] "WT"NOTE: lognormal distribution AIC/loglik/etc is on normal scale. Therefore, you can compare the AICs between fit.lnorm and fit.addProp since they are calculated on the same scale.
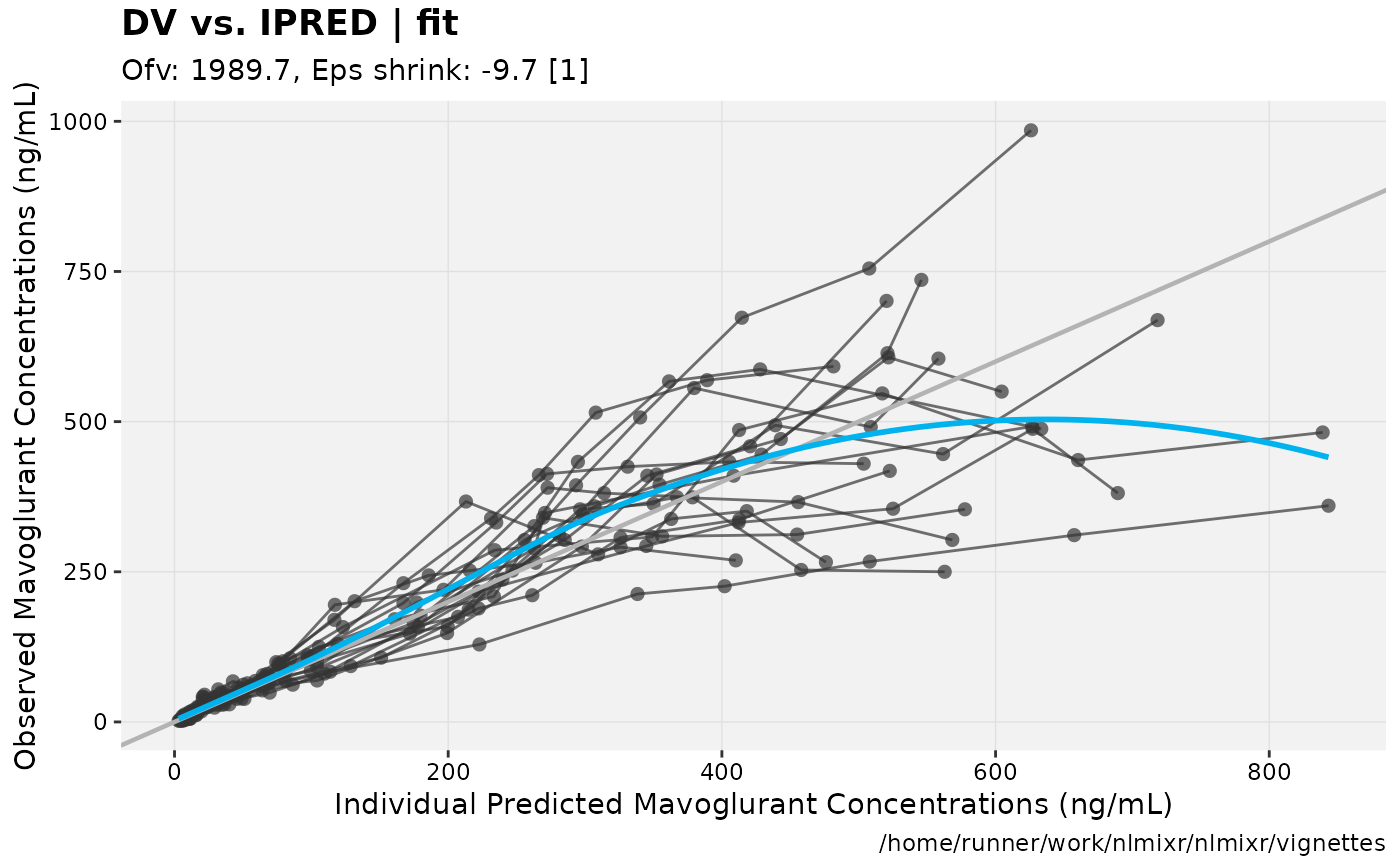
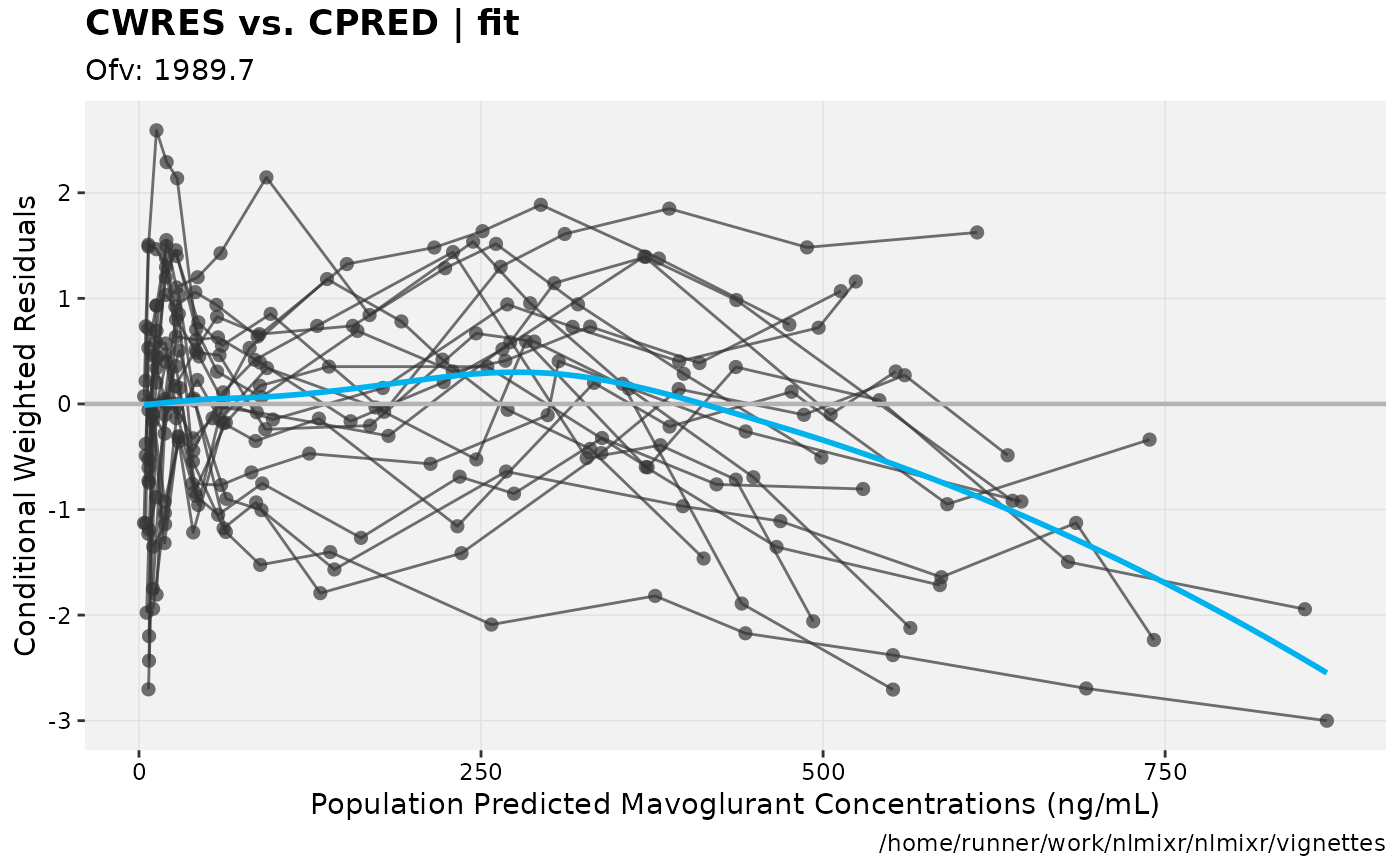
In this case you can see that the AIC for the log-normal model is better than the AIC for the addProp model.
gofs(fit.lnorm.S);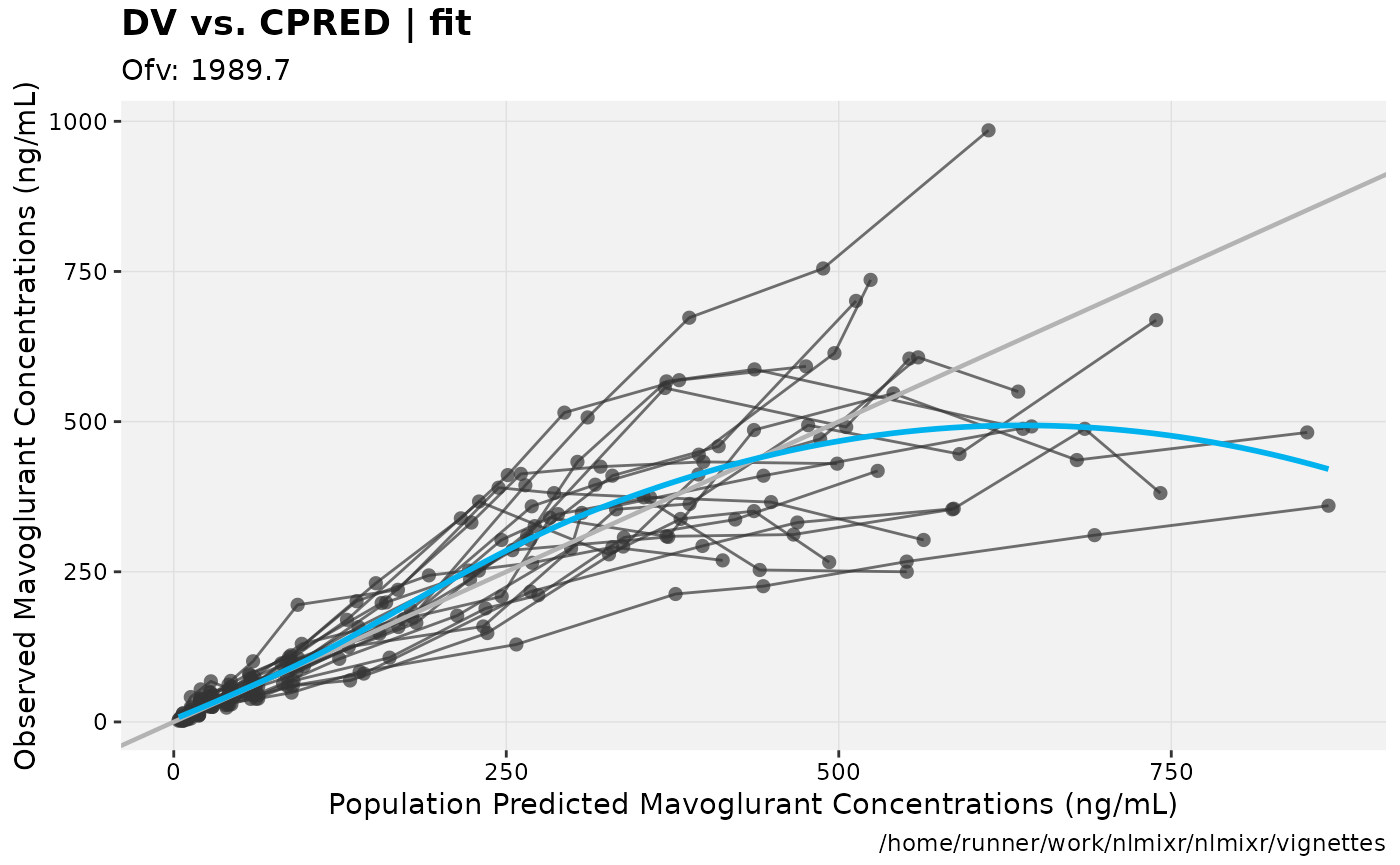
Piping to FOCEi
You can pipe models from different estimation methods to new estimation methods.
Additive + Proportional
fit.addProp.F <- fit.addProp.S %>%
nlmixr(est="focei",
control=list(print=0),
table=list(cwres=TRUE, npde=TRUE));
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [1] "WT"
#> calculating covariance matrix
#> [====|====|====|====|====|====|====|====|====|====] 0:01:38
#> done
## Since this was a model pipline, the data
## remains the same as the last fit.
gofs(fit.addProp.F);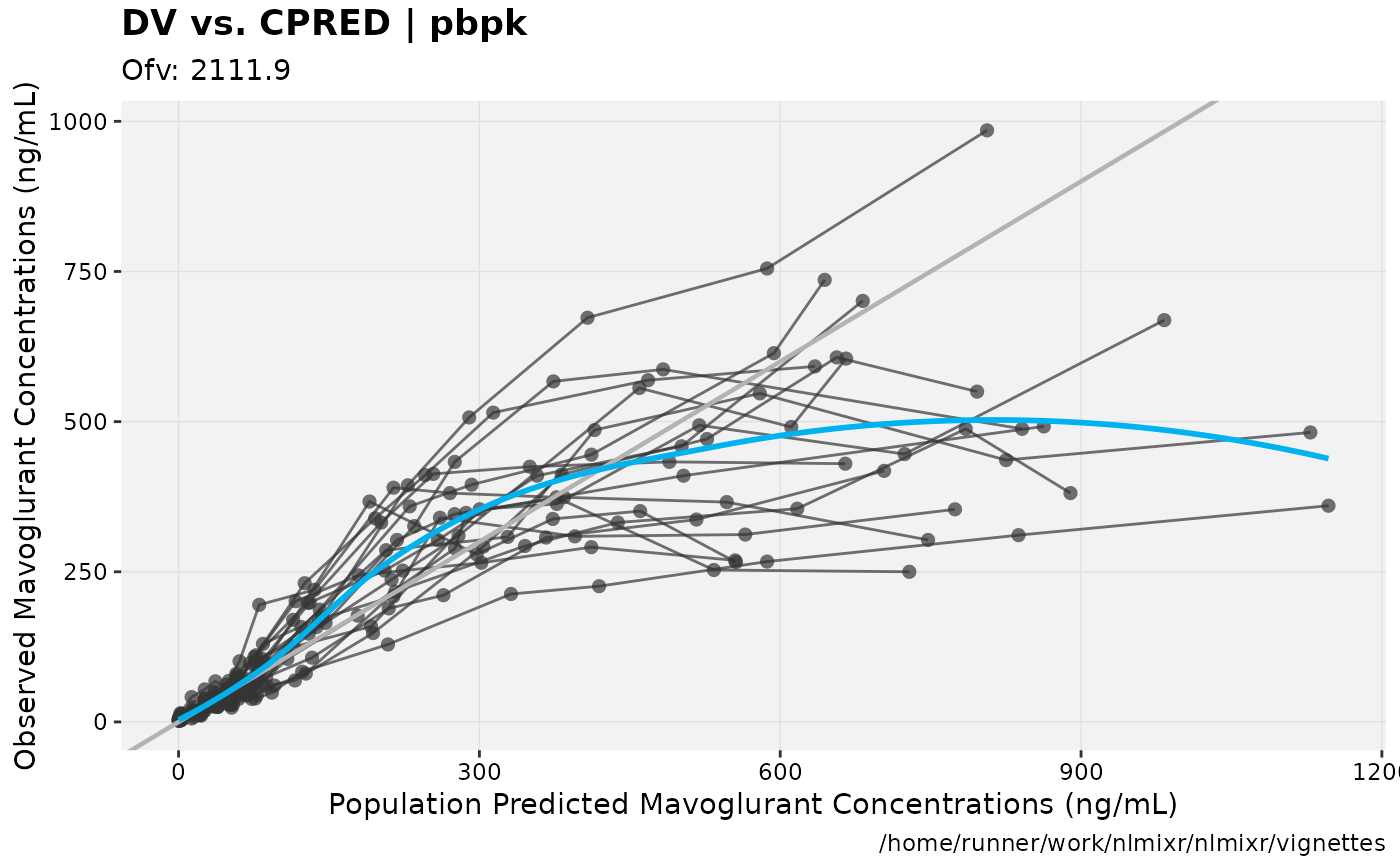
Lognoral
fit.lnorm.F <- fit.addProp.F %>%
model({C15 ~ lnorm(lnorm.err)}) %>%
nlmixr(est="focei",
control=list(print=0),
table=list(cwres=TRUE, npde=TRUE));
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [1] "WT"
#> calculating covariance matrix
#> [====|====|====|====|====|====|====|====|====|====] 0:00:20
#> done
## In this model pipline we are changing the fit method to focei.
gofs(fit.lnorm.F);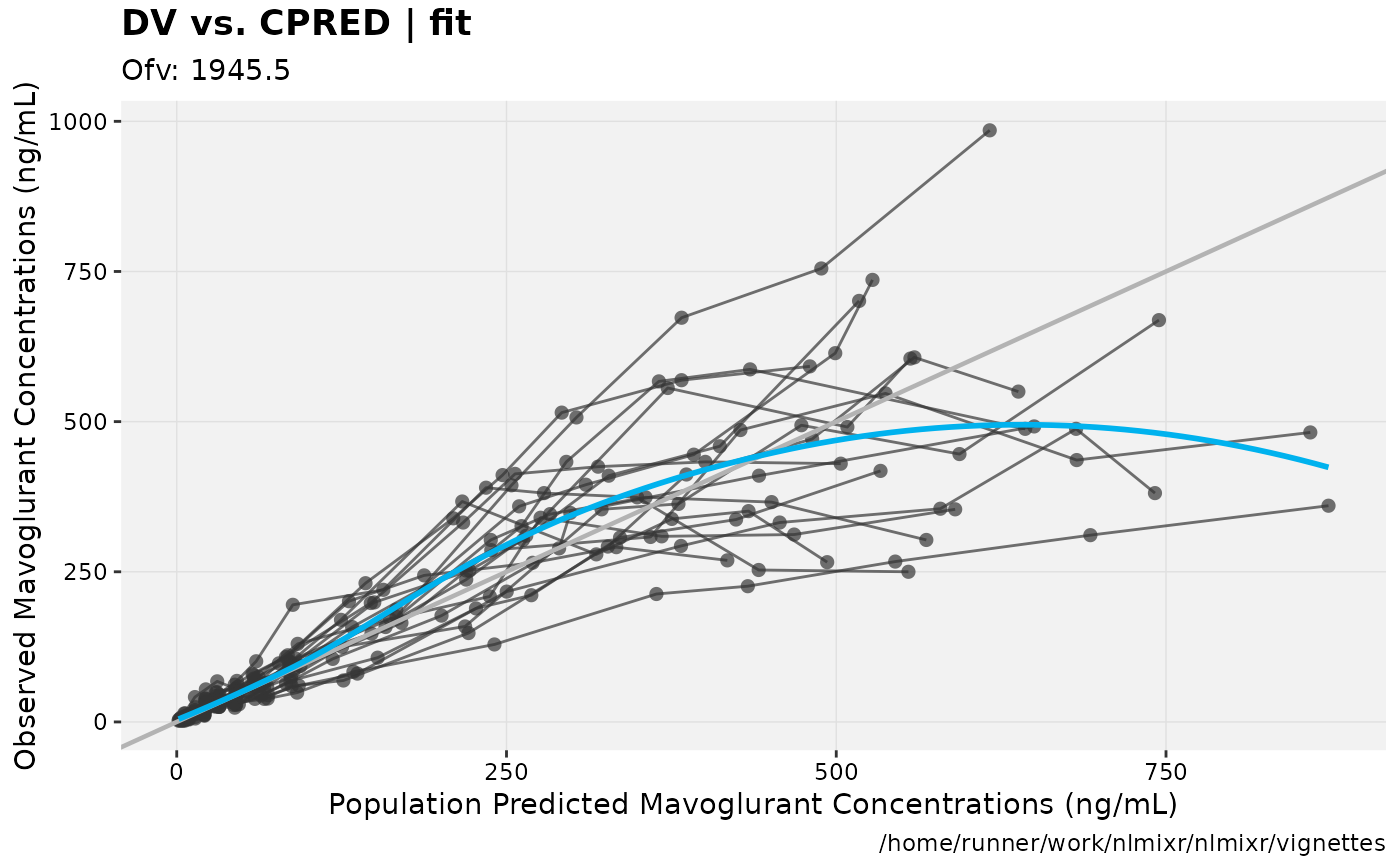
 # Traditional lognormal estimates are identical
# Traditional lognormal estimates are identical
datL <- dat
datL$DV <- log(datL$DV);
pbpkL <- function() {
ini({
##theta=exp(c(1.1, .3, 2, 7.6, .003, .3))
lKbBR = 1.1
lKbMU = 0.3
lKbAD = 2
lCLint = 7.6
lKbBO = 0.03
lKbRB = 0.3
eta.LClint ~ 4
add.err <- 1
})
model({
KbBR = exp(lKbBR)
KbMU = exp(lKbMU)
KbAD = exp(lKbAD)
CLint= exp(lCLint + eta.LClint)
KbBO = exp(lKbBO)
KbRB = exp(lKbRB)
## Regional blood flows
CO = (187.00*WT^0.81)*60/1000; # Cardiac output (L/h) from White et al (1968)
QHT = 4.0 *CO/100;
QBR = 12.0*CO/100;
QMU = 17.0*CO/100;
QAD = 5.0 *CO/100;
QSK = 5.0 *CO/100;
QSP = 3.0 *CO/100;
QPA = 1.0 *CO/100;
QLI = 25.5*CO/100;
QST = 1.0 *CO/100;
QGU = 14.0*CO/100;
QHA = QLI - (QSP + QPA + QST + QGU); # Hepatic artery blood flow
QBO = 5.0 *CO/100;
QKI = 19.0*CO/100;
QRB = CO - (QHT + QBR + QMU + QAD + QSK + QLI + QBO + QKI);
QLU = QHT + QBR + QMU + QAD + QSK + QLI + QBO + QKI + QRB;
## Organs' volumes = organs' weights / organs' density
VLU = (0.76 *WT/100)/1.051;
VHT = (0.47 *WT/100)/1.030;
VBR = (2.00 *WT/100)/1.036;
VMU = (40.00*WT/100)/1.041;
VAD = (21.42*WT/100)/0.916;
VSK = (3.71 *WT/100)/1.116;
VSP = (0.26 *WT/100)/1.054;
VPA = (0.14 *WT/100)/1.045;
VLI = (2.57 *WT/100)/1.040;
VST = (0.21 *WT/100)/1.050;
VGU = (1.44 *WT/100)/1.043;
VBO = (14.29*WT/100)/1.990;
VKI = (0.44 *WT/100)/1.050;
VAB = (2.81 *WT/100)/1.040;
VVB = (5.62 *WT/100)/1.040;
VRB = (3.86 *WT/100)/1.040;
## Fixed parameters
BP = 0.61; # Blood:plasma partition coefficient
fup = 0.028; # Fraction unbound in plasma
fub = fup/BP; # Fraction unbound in blood
KbLU = exp(0.8334);
KbHT = exp(1.1205);
KbSK = exp(-.5238);
KbSP = exp(0.3224);
KbPA = exp(0.3224);
KbLI = exp(1.7604);
KbST = exp(0.3224);
KbGU = exp(1.2026);
KbKI = exp(1.3171);
##-----------------------------------------
S15 = VVB*BP/1000;
C15 = Venous_Blood/S15
lnC15 = log(C15);
##-----------------------------------------
d/dt(Lungs) = QLU*(Venous_Blood/VVB - Lungs/KbLU/VLU);
d/dt(Heart) = QHT*(Arterial_Blood/VAB - Heart/KbHT/VHT);
d/dt(Brain) = QBR*(Arterial_Blood/VAB - Brain/KbBR/VBR);
d/dt(Muscles) = QMU*(Arterial_Blood/VAB - Muscles/KbMU/VMU);
d/dt(Adipose) = QAD*(Arterial_Blood/VAB - Adipose/KbAD/VAD);
d/dt(Skin) = QSK*(Arterial_Blood/VAB - Skin/KbSK/VSK);
d/dt(Spleen) = QSP*(Arterial_Blood/VAB - Spleen/KbSP/VSP);
d/dt(Pancreas) = QPA*(Arterial_Blood/VAB - Pancreas/KbPA/VPA);
d/dt(Liver) = QHA*Arterial_Blood/VAB + QSP*Spleen/KbSP/VSP + QPA*Pancreas/KbPA/VPA + QST*Stomach/KbST/VST + QGU*Gut/KbGU/VGU - CLint*fub*Liver/KbLI/VLI - QLI*Liver/KbLI/VLI;
d/dt(Stomach) = QST*(Arterial_Blood/VAB - Stomach/KbST/VST);
d/dt(Gut) = QGU*(Arterial_Blood/VAB - Gut/KbGU/VGU);
d/dt(Bones) = QBO*(Arterial_Blood/VAB - Bones/KbBO/VBO);
d/dt(Kidneys) = QKI*(Arterial_Blood/VAB - Kidneys/KbKI/VKI);
d/dt(Arterial_Blood) = QLU*(Lungs/KbLU/VLU - Arterial_Blood/VAB);
d/dt(Venous_Blood) = QHT*Heart/KbHT/VHT + QBR*Brain/KbBR/VBR +
QMU*Muscles/KbMU/VMU + QAD*Adipose/KbAD/VAD +
QSK*Skin/KbSK/VSK + QLI*Liver/KbLI/VLI + QBO*Bones/KbBO/VBO +
QKI*Kidneys/KbKI/VKI + QRB*Rest_of_Body/KbRB/VRB - QLU*Venous_Blood/VVB;
d/dt(Rest_of_Body) = QRB*(Arterial_Blood/VAB - Rest_of_Body/KbRB/VRB);
lnC15 ~ add(add.err)
})
}
fit.lnorm.trans <- pbpkL %>%
nlmixr(datL,est="saem",
control=list(print=0),
table=list(npde=TRUE, cwres=TRUE))
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [====|====|====|====|====|====|====|====|====|====] 0:00:00
#>
#> [1] "WT"NOTE: the estimates are the same but the AIC is different since it is calculated on the log scale.